Multiplex amplification enabled by selective circularization of large sets of genomic DNA fragments
- PMID: 15860768
- PMCID: PMC1087789
- DOI: 10.1093/nar/gni070
Multiplex amplification enabled by selective circularization of large sets of genomic DNA fragments
Abstract
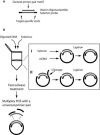
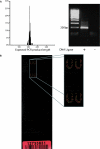
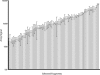
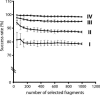
We present a method to specifically select large sets of DNA sequences for parallel amplification by PCR using target-specific oligonucleotide constructs, so-called selectors. The selectors are oligonucleotide duplexes with single-stranded target-complementary end-sequences that are linked by a general sequence motif. In the selection process, a pool of selectors is combined with denatured restriction digested DNA. Each selector hybridizes to its respective target, forming individual circular complexes that are covalently closed by enzymatic ligation. Non-circularized fragments are removed by exonucleolysis, enriching for the selected fragments. The general sequence that is introduced into the circularized fragments allows them to be amplified in parallel using a universal primer pair. The procedure avoids amplification artifacts associated with conventional multiplex PCR where two primers are used for each target, thereby reducing the number of amplification reactions needed for investigating large sets of DNA sequences. We demonstrate the specificity, reproducibility and flexibility of this process by performing a 96-plex amplification of an arbitrary set of specific DNA sequences, followed by hybridization to a cDNA microarray. Eighty-nine percent of the selectors generated PCR products that hybridized to the expected positions on the array, while little or no amplification artifacts were observed.
Figures
References
-
- Saiki R.K., Scharf S., Faloona F., Mullis K.B., Horn G.T., Erlich H.A., Arnheim N. Enzymatic amplification of beta-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science. 1985;230:1350–1354. - PubMed
-
- Telenius H., Carter N.P., Bebb C.E., Nordenskjold M., Ponder B.A., Tunnacliffe A. Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics. 1992;13:718–725. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

