A Kaposi's sarcoma virus RNA element that increases the nuclear abundance of intronless transcripts
- PMID: 15861127
- PMCID: PMC1142595
- DOI: 10.1038/sj.emboj.7600662
A Kaposi's sarcoma virus RNA element that increases the nuclear abundance of intronless transcripts
Abstract
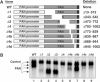
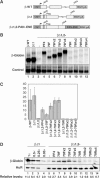
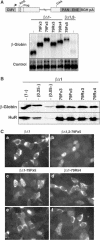
The Kaposi's sarcoma-associated herpesvirus produces a 1077 nucleotide noncoding, polyadenylated, exclusively nuclear RNA called PAN that is highly expressed in lytically infected cells. We report that PAN contains a novel post-transcriptional element essential for its abundant accumulation. The element, PAN-ENE (PAN RNA expression and nuclear retention element), increases the efficiency of 3'-end formation in vivo and is sufficient to enhance RNA abundance from an otherwise inefficiently expressed intronless beta-globin construct. The PAN-ENE does not concomitantly increase the production of encoded protein. Rather, it retains the unspliced beta-globin mRNA in the nucleus. Tethering of export factors can override the nuclear retention of the PAN-ENE, supporting a mechanism whereby the PAN-ENE blocks assembly of an export-competent mRNP. The activities of the PAN-ENE are specific to intronless constructs, since inserting the PAN-ENE into a spliced beta-globin construct has no effect on mRNA abundance and does not affect localization. This is the first characterization of a cis-acting element that increases RNA abundance of intronless transcripts but inhibits assembly of an export-competent mRNP.
Figures





References
-
- Andersen AA, Panning B (2003) Epigenetic gene regulation by noncoding RNAs. Curr Opin Cell Biol 15: 281–289 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

