Piv site-specific invertase requires a DEDD motif analogous to the catalytic center of the RuvC Holliday junction resolvases
- PMID: 15866929
- PMCID: PMC1112027
- DOI: 10.1128/JB.187.10.3431-3437.2005
Piv site-specific invertase requires a DEDD motif analogous to the catalytic center of the RuvC Holliday junction resolvases
Abstract
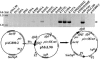
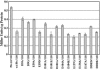
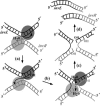
Piv, a unique prokaryotic site-specific DNA invertase, is related to transposases of the insertion elements from the IS110/IS492 family and shows no similarity to the site-specific recombinases of the tyrosine- or serine-recombinase families. Piv tertiary structure is predicted to include the RNase H-like fold that typically encompasses the catalytic site of the recombinases or nucleases of the retroviral integrase superfamily, including transposases and RuvC-like Holliday junction resolvases. Analogous to the DDE and DEDD catalytic motifs of transposases and RuvC, respectively, four Piv acidic residues D9, E59, D101, and D104 appear to be positioned appropriately within the RNase H fold to coordinate two divalent metal cations. This suggests mechanistic similarity between site-specific inversion mediated by Piv and transposition or endonucleolytic reactions catalyzed by enzymes of the retroviral integrase superfamily. The role of the DEDD motif in Piv catalytic activity was addressed using Piv variants that are substituted individually or multiply at these acidic residues and assaying for in vivo inversion, intermolecular recombination, and DNA binding activities. The results indicate that all four residues of the DEDD motif are required for Piv catalytic activity. The DEDD residues are not essential for inv recombination site recognition and binding, but this acidic tetrad does appear to contribute to the stability of Piv-inv interactions. On the basis of these results, a working model for Piv-mediated inversion that includes resolution of a Holliday junction is presented.
Figures
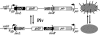
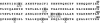



Similar articles
-
Conserved amino acid motifs from the novel Piv/MooV family of transposases and site-specific recombinases are required for catalysis of DNA inversion by Piv.Mol Microbiol. 2001 Feb;39(3):641-51. doi: 10.1046/j.1365-2958.2001.02276.x. Mol Microbiol. 2001. PMID: 11169105
-
A novel IS element, IS621, of the IS110/IS492 family transposes to a specific site in repetitive extragenic palindromic sequences in Escherichia coli.J Bacteriol. 2003 Aug;185(16):4891-900. doi: 10.1128/JB.185.16.4891-4900.2003. J Bacteriol. 2003. PMID: 12897009 Free PMC article.
-
Multiple DNA binding activities of the novel site-specific recombinase, Piv, from Moraxella lacunata.J Biol Chem. 1999 Apr 2;274(14):9698-706. doi: 10.1074/jbc.274.14.9698. J Biol Chem. 1999. PMID: 10092658
-
Transposition and site-specific recombination: adapting DNA cut-and-paste mechanisms to a variety of genetic rearrangements.FEMS Microbiol Rev. 1997 Sep;21(2):157-78. doi: 10.1111/j.1574-6976.1997.tb00349.x. FEMS Microbiol Rev. 1997. PMID: 9348666 Review.
-
Mechanisms of site-specific recombination.Annu Rev Biochem. 2006;75:567-605. doi: 10.1146/annurev.biochem.73.011303.073908. Annu Rev Biochem. 2006. PMID: 16756503 Review.
Cited by
-
A family of insertion sequences that impacts integrons by specific targeting of gene cassette recombination sites, the IS1111-attC Group.J Bacteriol. 2008 Jul;190(14):4959-70. doi: 10.1128/JB.00229-08. Epub 2008 May 16. J Bacteriol. 2008. PMID: 18487340 Free PMC article.
-
Site-specific insertion of IS492 in Pseudoalteromonas atlantica.J Bacteriol. 2009 Oct;191(20):6408-14. doi: 10.1128/JB.00771-09. Epub 2009 Aug 14. J Bacteriol. 2009. PMID: 19684137 Free PMC article.
-
Structural mechanism of bridge RNA-guided recombination.Nature. 2024 Jun;630(8018):994-1002. doi: 10.1038/s41586-024-07570-2. Epub 2024 Jun 26. Nature. 2024. PMID: 38926616 Free PMC article.
-
Plasticity of the P junc promoter of ISEc11, a new insertion sequence of the IS1111 family.J Bacteriol. 2006 Jul;188(13):4681-9. doi: 10.1128/JB.00332-06. J Bacteriol. 2006. PMID: 16788177 Free PMC article.
-
Integrating prokaryotes and eukaryotes: DNA transposases in light of structure.Crit Rev Biochem Mol Biol. 2010 Feb;45(1):50-69. doi: 10.3109/10409230903505596. Crit Rev Biochem Mol Biol. 2010. PMID: 20067338 Free PMC article. Review.
References
-
- Ariyoshi, M., D. G. Vassylyev, H. Iwasaki, H. Nakamura, H. Shinagawa, and K. Morikawa. 1994. Atomic structure of the RuvC resolvase: a Holliday junction-specific endonuclease from E. coli. Cell 78:1063-1072. - PubMed
-
- Ausubel, F. M., R. Brent, R. E. Kingston, D. D. Moore, J. G. Seidman, J. A. Smith, and K. Struhl. 1991. Current protocols in molecular biology. John Wiley & Sons, Inc., New York, N.Y.
-
- Bennett, R. J., H. J. Dunderdale, and S. C. West. 1993. Resolution of Holliday junctions by RuvC resolvase: cleavage specificity and DNA distortion. Cell 74:1021-1031. - PubMed
-
- Bennett, R. J., and S. C. West. 1995. Structural analysis of the RuvC-Holliday junction complex reveals an unfolded junction. J. Mol. Biol. 252:213-226. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

