Rapid and accurate identification of human isolates of Pasteurella and related species by sequencing the sodA gene
- PMID: 15872260
- PMCID: PMC1153776
- DOI: 10.1128/JCM.43.5.2307-2314.2005
Rapid and accurate identification of human isolates of Pasteurella and related species by sequencing the sodA gene
Abstract
The identification of Pasteurella and related bacteria remains a challenge. Here, a 449- to 473-bp fragment (sodA(int)) internal to the sodA gene, encoding the manganese-dependent superoxide dismutase, was amplified and sequenced with a single pair of degenerate primers from the type strains of Pasteurella (18 strains), Gallibacterium (1 strain), and Mannheimia (5 strains) species. The sodA(int)-based phylogenetic tree was in general agreement with that inferred from the analysis of the corresponding 16S rRNA gene sequences, with members of the Pasteurella sensu stricto cluster (Pasteurella multocida, Pasteurella canis, Pasteurella dagmatis, and Pasteurella stomatis) forming a monophyletic group and Gallibacterium and Mannheimia being independent monophyletic genera. However, the sodA(int) sequences showed a markedly higher divergence than the corresponding 16S rRNA genes, confirming that sodA is a potent target to differentiate related species. Thirty-three independent human clinical isolates phenotypically assigned to 13 Pasteurella species by a reference laboratory were successfully identified by comparing their sodA(int) sequences to those of the type species. In the course of this work, we identified the first Gallibacterium anatis isolate ever reported from a human clinical specimen. The sodA(int) sequences of the clinical isolates displayed less than 2.5% divergence from those of the corresponding type strains, except for the Pasteurella pneumotropica isolates, which were closely related to each other (> 98% sodA(int) sequence identity) but shared only 92% sodA(int) identity with the type strain. The method described here provides a rapid and accurate tool for species identification of Pasteurella isolates when access to a sequencing facility is available.
Figures

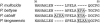

References
-
- Angen, Ø., R. Mutters, D. A. Caugant, J. E. Olsen, and M. Bisgaard. 1999. Taxonomic relationships of the [Pasteurella] haemolytica complex as evaluated by DNA-DNA hybridizations and 16S rRNA sequencing with proposal of Mannheimia haemolytica gen. nov., comb. nov., Mannheimia granulomatis comb. nov., Mannheimia glucosida sp. nov., Mannheimia ruminalis sp. nov. and Mannheimia varigena sp. nov. Int. J. Syst. Bacteriol. 49:67-86. - PubMed
-
- Birkbeck, T. H., L. A. Laidler, A. N. Grant, and D. I. Cox. 2002. Pasteurella skyensis sp. nov., isolated from Atlantic salmon (Salmo salar L.). Int. J. Syst. Evol. Microbiol. 52:699-704. - PubMed
-
- Bisgaard, M. 1993. Ecology and significance of Pasteurellaceae in animals. Zentralbl. Bakteriol. 279:7-26. - PubMed
-
- Catry, B., M. Baele, G. Opsomer, A. de Kruif, A. Decostere, and F. Haesebrouck. 2004. tRNA-intergenic spacer PCR for the identification of Pasteurella and Mannheimia spp. Vet. Microbiol. 98:251-260. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

