Brain lipid-binding protein is a direct target of Notch signaling in radial glial cells
- PMID: 15879553
- PMCID: PMC1091737
- DOI: 10.1101/gad.1302105
Brain lipid-binding protein is a direct target of Notch signaling in radial glial cells
Abstract
Radial glia function during CNS development both as neural progenitors and as a scaffolding supporting neuronal migration. To elucidate pathways involved in these functions, we mapped in vivo the promoter for Blbp, a radial glial gene. We show here that a binding site for the Notch effector CBF1 is essential for all Blbp transcription in radial glia, and that BLBP expression is significantly reduced in the forebrains of mice lacking the Notch1 and Notch3 receptors. These results identify Blbp as the first predominantly CNS-specific Notch target gene and suggest that it mediates some aspects of Notch signaling in radial glia.
Figures
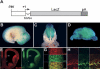
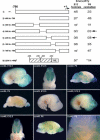


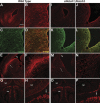
References
-
- Angevine J.B. and Sidman, R.L. 1961. Autoradiographic study of cell migration during histogenesis of cerebral cortex in the mouse. Nature 192: 766–768. - PubMed
-
- Anthony T.E., Klein, C., Fishell, G., and Heintz, N. 2004. Radial glia serve as neuronal progenitors in all regions of the central nervous system. Neuron 41: 881–890. - PubMed
-
- Anton E .S., Marchionni, M.A., Lee, K.F., and Rakic, P. 1997. Role of GGF/neuregulin signaling in interactions between migrating neurons and radial glia in the developing cerebral cortex. Development 124: 3501–3510. - PubMed
-
- Barolo S., Walker, R.G., Polyanovsky, A.D., Freschi, G., Keil, T., and Posakony, J.W. 2000. A notch-independent activity of suppressor of hairless is required for normal mechanoreceptor physiology. Cell 103: 957–969. - PubMed
-
- Fan X., Mikolaenko, I., Elhassan, I., Ni, X., Wang, Y., Ball, D., Brat, D.J., Perry, A., and Eberhart, C.G. 2004. Notch1 and notch2 have opposite effects on embryonal brain tumor growth. Cancer Res. 64: 7787–7793. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
