Conservation and evolvability in regulatory networks: the evolution of ribosomal regulation in yeast
- PMID: 15883364
- PMCID: PMC1091753
- DOI: 10.1073/pnas.0502521102
Conservation and evolvability in regulatory networks: the evolution of ribosomal regulation in yeast
Abstract
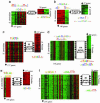
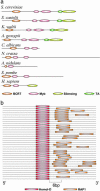
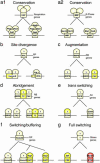
Transcriptional modules of coregulated genes play a key role in regulatory networks. Comparative studies show that modules of coexpressed genes are conserved across taxa. However, little is known about the mechanisms underlying the evolution of module regulation. Here, we explore the evolution of cis-regulatory programs associated with conserved modules by integrating expression profiles for two yeast species and sequence data for a total of 17 fungal genomes. We show that although the cis-elements accompanying certain conserved modules are strictly conserved, those of other conserved modules are remarkably diverged. In particular, we infer the evolutionary history of the regulatory program governing ribosomal modules. We show how a cis-element emerged concurrently in dozens of promoters of ribosomal protein genes, followed by the loss of a more ancient cis-element. We suggest that this formation of an intermediate redundant regulatory program allows conserved transcriptional modules to gradually switch from one regulatory mechanism to another while maintaining their functionality. Our work provides a general framework for the study of the dynamics of promoter evolution at the level of transcriptional modules and may help in understanding the evolvability and increased redundancy of transcriptional regulation in higher organisms.
Figures

References
-
- Ihmels, J., Friedlander, G., Bergmann, S., Sarig, O., Ziv, Y. & Barkai, N. (2002) Nat. Genet. 31, 370-377. - PubMed
-
- Segal, E., Shapira, M., Regev, A., Pe'er, D., Botstein, D., Koller, D. & Friedman, N. (2003) Nat. Genet. 34, 166-176. - PubMed
-
- Stuart, J. M., Segal, E., Koller, D. & Kim, S. K. (2003) Science 302, 249-255. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

