Large-scale identification of expressed sequence tags involved in rice and rice blast fungus interaction
- PMID: 15888683
- PMCID: PMC1104166
- DOI: 10.1104/pp.104.055624
Large-scale identification of expressed sequence tags involved in rice and rice blast fungus interaction
Abstract
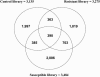
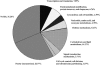
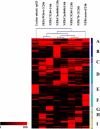
To better understand the molecular basis of the defense response against the rice blast fungus (Magnaporthe grisea), a large-scale expressed sequence tag (EST) sequencing approach was used to identify genes involved in the early infection stages in rice (Oryza sativa). Six cDNA libraries were constructed using infected leaf tissues harvested from 6 conditions: resistant, partially resistant, and susceptible reactions at both 6 and 24 h after inoculation. Two additional libraries were constructed using uninoculated leaves and leaves from the lesion mimic mutant spl11. A total of 68,920 ESTs were generated from 8 libraries. Clustering and assembly analyses resulted in 13,570 unique sequences from 10,934 contigs and 2,636 singletons. Gene function classification showed that 42% of the ESTs were predicted to have putative gene function. Comparison of the pathogen-challenged libraries with the uninoculated control library revealed an increase in the percentage of genes in the functional categories of defense and signal transduction mechanisms and cell cycle control, cell division, and chromosome partitioning. In addition, hierarchical clustering analysis grouped the eight libraries based on their disease reactions. A total of 7,748 new and unique ESTs were identified from our collection compared with the KOME full-length cDNA collection. Interestingly, we found that rice ESTs are more closely related to sorghum (Sorghum bicolor) ESTs than to barley (Hordeum vulgare), wheat (Triticum aestivum), and maize (Zea mays) ESTs. The large cataloged collection of rice ESTs in this study provides a solid foundation for further characterization of the rice defense response and is a useful public genomic resource for rice functional genomics studies.
Figures

References
-
- Adams M, Kerlavage AR, Fleischmann RD, Fuldner RA, Bult CJ, Lee NH, Kirkness EF, Weinstock KG, Gocayne JD, White O, et al (1995) Initial assessment of human gene diversity and expression patterns based upon 83 million nucleotides of cDNA sequence. Nature 377: 173–174 - PubMed
-
- Becraft PW (2002) Receptor kinase signaling in plant development. Annu Rev Cell Dev Biol 18: 163–192 - PubMed
-
- Birch PRJ, Kamoun S (2000) Studying interaction transcriptomes: coordinated analyses of gene expression during plant-microorganism interactions. In R Wood, ed, New Technologies for Life Sciences: A Trends Guide. Elsevier Science, New York, pp 77–82
-
- Brenner S, Johnson M, Bridgham J, Golda G, Lloyd DH, Johnson D, Luo S, McCurdy S, Foy M, Ewan M, et al (2000) Gene expression analysis by massively parallel signature sequencing (MPSS) on microbead arrays. Nat Biotechnol 18: 630–634 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

