The cellular chaperone heat shock protein 90 facilitates Flock House virus RNA replication in Drosophila cells
- PMID: 15890922
- PMCID: PMC1112161
- DOI: 10.1128/JVI.79.11.6827-6837.2005
The cellular chaperone heat shock protein 90 facilitates Flock House virus RNA replication in Drosophila cells
Abstract
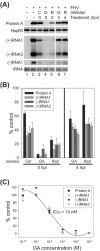
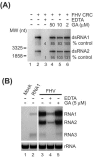
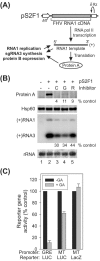
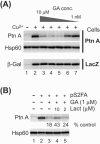
The assembly of viral RNA replication complexes on intracellular membranes represents a critical step in the life cycle of positive-strand RNA viruses. We investigated the role of the cellular chaperone heat shock protein 90 (Hsp90) in viral RNA replication complex assembly and function using Flock House virus (FHV), an alphanodavirus whose RNA-dependent RNA polymerase, protein A, is essential for viral RNA replication complex assembly on mitochondrial outer membranes. The Hsp90 chaperone complex transports cellular mitochondrial proteins to the outer mitochondrial membrane import receptors, and thus we hypothesized that Hsp90 may also facilitate FHV RNA replication complex assembly or function. Treatment of FHV-infected Drosophila S2 cells with the Hsp90-specific inhibitor geldanamycin or radicicol potently suppressed the production of infectious virions and the accumulation of protein A and genomic, subgenomic, and template viral RNA. In contrast, geldanamycin did not inhibit the activity of preformed FHV RNA replication complexes. Hsp90 inhibitors also suppressed viral RNA and protein A accumulation in S2 cells expressing an FHV RNA replicon. Furthermore, Hsp90 inhibition with either geldanamycin or RNAi-mediated chaperone downregulation suppressed protein A accumulation in the absence of viral RNA replication. These results identify Hsp90 as a host factor involved in FHV RNA replication and suggest that FHV uses established cellular chaperone pathways to assemble its RNA replication complexes on intracellular membranes.
Figures

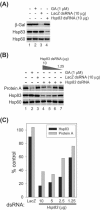
Similar articles
-
A functional heat shock protein 90 chaperone is essential for efficient flock house virus RNA polymerase synthesis in Drosophila cells.J Virol. 2007 Aug;81(16):8412-20. doi: 10.1128/JVI.00189-07. Epub 2007 May 23. J Virol. 2007. PMID: 17522196 Free PMC article.
-
The heat shock protein 70 cochaperone YDJ1 is required for efficient membrane-specific flock house virus RNA replication complex assembly and function in Saccharomyces cerevisiae.J Virol. 2008 Feb;82(4):2004-12. doi: 10.1128/JVI.02017-07. Epub 2007 Dec 5. J Virol. 2008. PMID: 18057252 Free PMC article.
-
Flock house virus RNA replicates on outer mitochondrial membranes in Drosophila cells.J Virol. 2001 Dec;75(23):11664-76. doi: 10.1128/JVI.75.23.11664-11676.2001. J Virol. 2001. PMID: 11689648 Free PMC article.
-
Recent insights into the biology and biomedical applications of Flock House virus.Cell Mol Life Sci. 2008 Sep;65(17):2675-87. doi: 10.1007/s00018-008-8037-y. Cell Mol Life Sci. 2008. PMID: 18516498 Free PMC article. Review.
-
An Insight to Heat Shock Protein 90: A Remedy for Multiple Problems.Curr Pharm Des. 2022;28(32):2664-2676. doi: 10.2174/1381612828666220829120630. Curr Pharm Des. 2022. PMID: 36043709 Review.
Cited by
-
Complementary transcriptomic, lipidomic, and targeted functional genetic analyses in cultured Drosophila cells highlight the role of glycerophospholipid metabolism in Flock House virus RNA replication.BMC Genomics. 2010 Mar 17;11:183. doi: 10.1186/1471-2164-11-183. BMC Genomics. 2010. PMID: 20236518 Free PMC article.
-
Human butyrate-induced transcript 1 interacts with hepatitis C virus NS5A and regulates viral replication.J Virol. 2008 Mar;82(6):2631-41. doi: 10.1128/JVI.02153-07. Epub 2007 Dec 26. J Virol. 2008. PMID: 18160438 Free PMC article.
-
Theiler's murine encephalomyelitis virus infection induces a redistribution of heat shock proteins 70 and 90 in BHK-21 cells, and is inhibited by novobiocin and geldanamycin.Cell Stress Chaperones. 2011 Sep;16(5):505-15. doi: 10.1007/s12192-011-0262-x. Epub 2011 Mar 29. Cell Stress Chaperones. 2011. PMID: 21445704 Free PMC article.
-
Maturation-dependent responses of human neuronal cells to western equine encephalitis virus infection and type I interferons.Virology. 2008 Mar 1;372(1):208-20. doi: 10.1016/j.virol.2007.10.025. Epub 2007 Nov 26. Virology. 2008. PMID: 18022665 Free PMC article.
-
Membrane-bound tomato mosaic virus replication proteins participate in RNA synthesis and are associated with host proteins in a pattern distinct from those that are not membrane bound.J Virol. 2006 Sep;80(17):8459-68. doi: 10.1128/JVI.00545-06. J Virol. 2006. PMID: 16912296 Free PMC article.
References
-
- Arbeitman, M. N., and D. S. Hogness. 2000. Molecular chaperones activate the Drosophila ecdysone receptor, an RXR heterodimer. Cell 101:67-77. - PubMed
-
- Ball, L. A., and K. L. Johnson. 1998. Nodaviruses of insects, p. 225-267. In L. K. Miller and L. A. Ball (ed.), The insect viruses. Plenum Publishing Corporation, New York, N.Y.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

