Structural basis for lysidine formation by ATP pyrophosphatase accompanied by a lysine-specific loop and a tRNA-recognition domain
- PMID: 15894617
- PMCID: PMC1140429
- DOI: 10.1073/pnas.0501003102
Structural basis for lysidine formation by ATP pyrophosphatase accompanied by a lysine-specific loop and a tRNA-recognition domain
Abstract
Lysidine, a lysine-combined modified cytidine, is exclusively located at the anticodon wobble position (position 34) of eubacterial tRNA(Ile)(2) and not only converts the codon specificity from AUG to AUA, but also converts the aminoacylation specificity from recognition by methionyl-tRNA synthetase to that by isoleucyl-tRNA synthetase (IleRS). Here, we report the crystal structure of lysidine synthetase (TilS) from Aquifex aeolicus at 2.42-A resolution. TilS forms a homodimer, and each subunit consists of the N-terminal dinucleotide-binding fold domain (NTD), with a characteristic central hole, and the C-terminal globular domain (CTD) connected by a long alpha-helical linker. The NTD shares striking structural similarity with the ATP-pyrophosphatase domain of GMP synthetase, which reminds us of the two-step reaction by TilS: adenylation of C34 and lysine attack on the C2 carbon. Conserved amino acid residues are clustered around the NTD central hole. Kinetic analyses of the conserved residues' mutants indicated that C34 of tRNA(Ile)(2) is adenylated by an ATP lying across the NTD central hole and that a lysine, which is activated at a loop appended to the NTD, nucleophilically attacks the C2 carbon from the rear. Escherichia coli TilS (called MesJ) has an additional CTD, which may recognize the tRNA(Ile)(2) acceptor stem. In contrast, a mutational study revealed that A. aeolicus TilS does not recognize the tRNA acceptor stem but recognizes the C29.G41 base pair in the anticodon stem. Thus, the two TilS enzymes discriminate tRNA(Ile)(2) from tRNA(Met) by strategies similar to that used by IleRS, but in distinct manners.
Figures

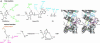
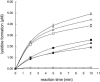
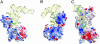
Similar articles
-
Discovery and characterization of tRNAIle lysidine synthetase (TilS).FEBS Lett. 2010 Jan 21;584(2):272-7. doi: 10.1016/j.febslet.2009.11.085. FEBS Lett. 2010. PMID: 19944692 Review.
-
molecular mechanism of lysidine synthesis that determines tRNA identity and codon recognition.Mol Cell. 2005 Jul 22;19(2):235-46. doi: 10.1016/j.molcel.2005.06.007. Mol Cell. 2005. PMID: 16039592
-
Structural basis for translational fidelity ensured by transfer RNA lysidine synthetase.Nature. 2009 Oct 22;461(7267):1144-8. doi: 10.1038/nature08474. Nature. 2009. PMID: 19847269
-
Enzymatic conversion of cytidine to lysidine in anticodon of bacterial isoleucyl-tRNA--an alternative way of RNA editing.Trends Biochem Sci. 2004 Apr;29(4):165-8. doi: 10.1016/j.tibs.2004.02.009. Trends Biochem Sci. 2004. PMID: 15124629 Review. No abstract available.
-
Mechanisms of the tRNA wobble cytidine modification essential for AUA codon decoding in prokaryotes.Biosci Biotechnol Biochem. 2015;79(3):347-53. doi: 10.1080/09168451.2014.975185. Epub 2014 Oct 28. Biosci Biotechnol Biochem. 2015. PMID: 25348586 Review.
Cited by
-
The C-terminal end of the Trypanosoma brucei editing deaminase plays a critical role in tRNA binding.RNA. 2011 Jul;17(7):1296-306. doi: 10.1261/rna.2748211. Epub 2011 May 20. RNA. 2011. PMID: 21602302 Free PMC article.
-
Transfer RNA Modification Enzymes from Thermophiles and Their Modified Nucleosides in tRNA.Microorganisms. 2018 Oct 20;6(4):110. doi: 10.3390/microorganisms6040110. Microorganisms. 2018. PMID: 30347855 Free PMC article. Review.
-
The ATPase activity of the G2alt gene encoding an aluminium tolerance protein from Anoxybacillus gonensis G2.J Microbiol. 2011 Aug;49(4):641-50. doi: 10.1007/s12275-011-0522-6. Epub 2011 Sep 2. J Microbiol. 2011. PMID: 21887649
-
Extensive import of nucleus-encoded tRNAs into chloroplasts of the photosynthetic lycophyte, Selaginella kraussiana.Proc Natl Acad Sci U S A. 2024 Nov 12;121(46):e2412221121. doi: 10.1073/pnas.2412221121. Epub 2024 Nov 6. Proc Natl Acad Sci U S A. 2024. PMID: 39503889 Free PMC article.
-
Differential annotation of tRNA genes with anticodon CAT in bacterial genomes.Nucleic Acids Res. 2006;34(20):6015-22. doi: 10.1093/nar/gkl739. Epub 2006 Oct 27. Nucleic Acids Res. 2006. PMID: 17071718 Free PMC article.
References
-
- Bjork, G. (1995) tRNA: Structure, Biosynthesis, and Function, eds. Soll, D. & RajBhandary, U. L. (American Society for Microbiology, Washington, D.C.)
-
- Schimmel, P. R. (1979) in Transfer RNA: Structure, Properties, and Recognition, eds. Schimmel, P. R., Soll, D. & Abelson, J. N. (Cold Spring Harbor Lab. Press, Plainview, NY), pp. 297–310.
-
- Ozeki, H., Inokuchi, H., Yamao, F., Kodaira, M., Sakano, H., Ikemura, T. & Shimura, Y. (1980) in Transfer RNA: Biological Aspects, eds. Soll, D., Abelson, J. N. & Schimmel, P. R. (Cold Spring Harbor Lab. Press, Plainview, NY), pp. 341–362.
-
- Gerber, A. P. & Keller, W. (1999) Science 286, 1146–1149. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

