Basic mechanism of eukaryotic chromosome segregation
- PMID: 15897183
- PMCID: PMC1569467
- DOI: 10.1098/rstb.2004.1615
Basic mechanism of eukaryotic chromosome segregation
Abstract
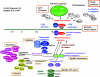
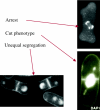
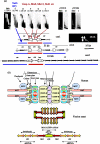
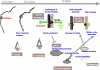
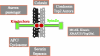
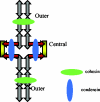
We now have firm evidence that the basic mechanism of chromosome segregation is similar among diverse eukaryotes as the same genes are employed. Even in prokaryotes, the very basic feature of chromosome segregation has similarities to that of eukaryotes. Many aspects of chromosome segregation are closely related to a cell cycle control that includes stage-specific protein modification and proteolysis. Destruction of mitotic cyclin and securin leads to mitotic exit and separase activation, respectively. Key players in chromosome segregation are SMC-containing cohesin and condensin, DNA topoisomerase II, APC/C ubiquitin ligase, securin-separase complex, aurora passengers, and kinetochore microtubule destabilizers or regulators. In addition, the formation of mitotic kinetochore and spindle apparatus is absolutely essential. The roles of principal players in basic chromosome segregation are discussed: most players have interphase as well as mitotic functions. A view on how the centromere/kinetochore is formed is described.
Figures

References
-
- Aono N, Sutani T, Tomonaga T, Mochida S, Yanagida M. Cnd2 has dual roles in mitotic condensation and interphase. Nature. 2002;417:197–202. - PubMed
-
- Axton J.M, Dombradi V, Cohen P.T.W, Glover D.M. One of the protein phosphatase 1 isozymes in Drosophila is essential for mitosis. Cell. 1990;63:33–46. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
