Iron-responsive regulation of the Helicobacter pylori iron-cofactored superoxide dismutase SodB is mediated by Fur
- PMID: 15901691
- PMCID: PMC1112043
- DOI: 10.1128/JB.187.11.3687-3692.2005
Iron-responsive regulation of the Helicobacter pylori iron-cofactored superoxide dismutase SodB is mediated by Fur
Abstract
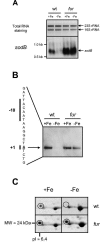
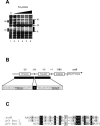
Maintaining iron homeostasis is a necessity for all living organisms, as free iron augments the generation of reactive oxygen species like superoxide anions, at the risk of subsequent lethal cellular damage. The iron-responsive regulator Fur controls iron metabolism in many bacteria, including the important human pathogen Helicobacter pylori, and thus is directly or indirectly involved in regulation of oxidative stress defense. Here we demonstrate that Fur is a direct regulator of the H. pylori iron-cofactored superoxide dismutase SodB, which is essential for the defense against toxic superoxide radicals. Transcription of the sodB gene was iron induced in H. pylori wild-type strain 26695, resulting in expression of the SodB protein in iron-replete conditions but an absence of expression in iron-restricted conditions. Mutation of the fur gene resulted in constitutive, iron-independent expression of SodB. Recombinant H. pylori Fur protein bound with low affinity to the sodB promoter region, but addition of the iron substitute Mn2+ abolished binding. The operator sequence of the iron-free form of Fur, as identified by DNase I footprinting, was located directly upstream of the sodB gene at positions -5 to -47 from the transcription start site. The direct role of Fur in regulation of the H. pylori sodB gene contrasts with the small-RNA-mediated sodB regulation observed in Escherichia coli. In conclusion, H. pylori Fur is a versatile regulator involved in many pathways essential for gastric colonization, including superoxide stress defense.
Figures

Similar articles
-
A single nucleotide change affects fur-dependent regulation of sodB in H. pylori.PLoS One. 2009;4(4):e5369. doi: 10.1371/journal.pone.0005369. Epub 2009 Apr 28. PLoS One. 2009. PMID: 19399190 Free PMC article.
-
Fur positive regulation of iron superoxide dismutase in Escherichia coli: functional analysis of the sodB promoter.J Bacteriol. 2000 Jul;182(13):3802-8. doi: 10.1128/JB.182.13.3802-3808.2000. J Bacteriol. 2000. PMID: 10850997 Free PMC article.
-
Two amino acids mutation of ferric uptake regulator determines Helicobacter pylori resistance to metronidazole.Antioxid Redox Signal. 2011 Jan 1;14(1):15-23. doi: 10.1089/ars.2010.3146. Epub 2010 Aug 26. Antioxid Redox Signal. 2011. PMID: 20518707
-
The ferric uptake regulator of Helicobacter pylori: a critical player in the battle for iron and colonization of the stomach.Future Microbiol. 2013 Jun;8(6):725-38. doi: 10.2217/fmb.13.43. Future Microbiol. 2013. PMID: 23701330 Free PMC article. Review.
-
[Study of infection strategies of Helicobacter pylori and host cell response against CagA oncoprotein].Nihon Saikingaku Zasshi. 2014;69(4):565-75. doi: 10.3412/jsb.69.565. Nihon Saikingaku Zasshi. 2014. PMID: 25447982 Review. Japanese.
Cited by
-
The nickel-responsive regulator NikR controls activation and repression of gene transcription in Helicobacter pylori.Infect Immun. 2005 Nov;73(11):7252-8. doi: 10.1128/IAI.73.11.7252-7258.2005. Infect Immun. 2005. PMID: 16239520 Free PMC article.
-
Pathogenesis of Helicobacter pylori infection.Clin Microbiol Rev. 2006 Jul;19(3):449-90. doi: 10.1128/CMR.00054-05. Clin Microbiol Rev. 2006. PMID: 16847081 Free PMC article. Review.
-
Integrative computational protocol for the discovery of inhibitors of the Helicobacter pylori nickel response regulator (NikR).J Mol Model. 2011 Dec;17(12):3075-84. doi: 10.1007/s00894-011-0962-2. Epub 2011 Mar 1. J Mol Model. 2011. PMID: 21360181
-
Snapshot of iron response in Shewanella oneidensis by gene network reconstruction.BMC Genomics. 2009 Mar 25;10:131. doi: 10.1186/1471-2164-10-131. BMC Genomics. 2009. PMID: 19321007 Free PMC article.
-
Transcription of IVIAT and Virulence Genes in Photobacterium damselae Subsp. piscicida Infecting Solea senegalensis.Microorganisms. 2018 Jul 12;6(3):67. doi: 10.3390/microorganisms6030067. Microorganisms. 2018. PMID: 30002314 Free PMC article.
References
-
- Alamuri, P., and R. J. Maier. 2005. Methionine sulphoxide reductase is an important antioxidant enzyme in the gastric pathogen Helicobacter pylori. Mol. Microbiol. 55:653. - PubMed
-
- Andrews, S. C., A. K. Robinson, and F. Rodriguez-Quinones. 2003. Bacterial iron homeostasis. FEMS Microbiol. Rev. 27:215-237. - PubMed
-
- Barnard, F. M., M. F. Loughlin, H. P. Fainberg, M. P. Messenger, D. W. Ussery, P. Williams, and P. J. Jenks. 2004. Global regulation of virulence and the stress response by CsrA in the highly adapted human gastric pathogen Helicobacter pylori. Mol. Microbiol. 51:15-32. - PubMed
-
- Bernhardt, J., K. Buttner, C. Scharf, and M. Hecker. 1999. Dual channel imaging of two-dimensional electropherograms in Bacillus subtilis. Electrophoresis 20:2225-2240. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

