Loss-of-function mutations of ROOT HAIR DEFECTIVE3 suppress root waving, skewing, and epidermal cell file rotation in Arabidopsis
- PMID: 15908600
- PMCID: PMC1150390
- DOI: 10.1104/pp.105.059774
Loss-of-function mutations of ROOT HAIR DEFECTIVE3 suppress root waving, skewing, and epidermal cell file rotation in Arabidopsis
Abstract
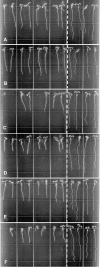
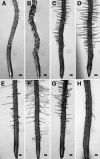
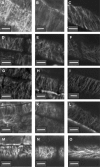
Wild-type Arabidopsis (Arabidopsis thaliana L. Heynh.) roots growing on a tilted surface of impenetrable hard-agar media adopt a wave-like pattern and tend to skew to the right of the gravity vector (when viewed from the back of the plate through the medium). Reversible root-tip rotation often accompanies the clockwise and counterclockwise curves that form each wave. These rotations are manifested by epidermal cell file rotation (CFR) along the root. Loss-of-function alleles of ROOT HAIR DEFECTIVE3 (RHD3), a gene previously implicated in the control of vesicle trafficking between the endoplasmic reticulum and the Golgi compartments, resulted in an almost complete suppression of epidermal CFR, root skewing, and waving on hard-agar surfaces. Several other root hair defective mutants (rhd2-1, rhd4-1, and rhd6-1) did not exhibit dramatic alterations in these root growth behaviors, suggesting that a generalized defect in root hair formation is not responsible for the surface-dependent phenotypes of rhd3. However, similar alterations in root growth behavior were observed in a variety of mutants characterized by defects in cell expansion (cob-1, cob-2, eto1-1, eto2-1, erh2-1, and erh3-1). The erh2-1 and rhd3-1 mutants differed from other anisotropic cell expansion mutants, though, by an inability to respond to low doses of the microtubule-binding drug propyzamide, which normally causes enhanced left-handed CFR and right skewing. We hypothesize that RHD3 may control epidermal CFR, root skewing, and waving on hard-agar surfaces by regulating the traffic of wall- or plasma membrane-associated determinants of anisotropic cell expansion.
Figures





References
-
- Andeme-Onzighi C, Sivaguru M, Judy-March J, Baskin T, Driouich A (2002) The reb1-1 mutation of Arabidopsis alters the morphology of trichoblasts, the expression of arabinogalactan-proteins and the organization of cortical microtubules. Planta 215: 949–958 - PubMed
-
- Ausubel F, Brent R, Kingston R, Moore D, Seidman J, Smith J, Struhl K (1994) Current Protcols in Molecular Biology—Updates 1994-2002. John Wiley and Sons, New York
-
- Baskin T, Betzner A, Hoggart R, Cork A, Williamson R (1992) Root morphology mutants in Arabidopsis thaliana. Aust J Plant Physiol 19: 427–437
-
- Baskin T, Wilson J, Cork A, Williamson R (1994) Morphology and microtubule organization in Arabidopsis roots exposed to oryzalin or taxol. Plant Cell Physiol 35: 935–942 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

