New NTP analogs: the synthesis of 4'-thioUTP and 4'-thioCTP and their utility for SELEX
- PMID: 15914669
- PMCID: PMC1140078
- DOI: 10.1093/nar/gki578
New NTP analogs: the synthesis of 4'-thioUTP and 4'-thioCTP and their utility for SELEX
Abstract
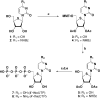
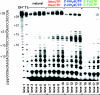
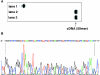
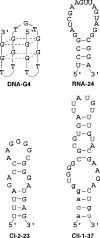
The synthesis of the triphosphates of 4'-thiouridine and 4'-thiocytidine, 4'-thioUTP (7; thioUTP) and 4'-thioCTP (8; thioCTP), and their utility for SELEX (systematic evolution of ligands by exponential enrichment) is described. The new nucleoside triphosphate (NTP) analogs 7 and 8 were prepared from appropriately protected 4'-thiouridine and -cytidine derivatives using the one-pot method reported by J. Ludwig and F. Eckstein [(1989) J. Org. Chem., 54, 631-635]. Because SELEX requires both in vitro transcription and reverse transcription, we examined the ability of 7 and 8 for SELEX by focusing on the two steps. Incorporation of 7 and 8 by T7 RNA polymerase to give 4'-thioRNA (thioRNA) proceeded well and was superior to those of the two sets of frequently used modified NTP analogs for SELEX (2'-NH2dUTP and 2'-NH2dCTP; 2'-FdUTP and 2'-FdCTP), when an adequate leader sequence of DNA template was selected. We revealed that a leader sequence of about +15 of DNA template is important for the effective incorporation of modified NTP analogs by T7 RNA polymerase. In addition, reverse transcription of the resulting thioRNA into the complementary DNA in the presence of 2'-deoxynucleoside triphosphates (dNTPs) also proceeded smoothly and precisely. The stability of the thioRNA toward RNase A was 50 times greater than that of the corresponding natural RNA. With these successful results in hand, we attempted the selection of thioRNA aptamers to human alpha-thrombin using thioUTP and thioCTP, and found a thioRNA aptamer with high binding affinity (K(d) = 4.7 nM).
Figures






Similar articles
-
In vitro selection of human alpha-thrombin RNA aptamer using 4'-thioUTP and 4'-thioCTP.Nucleic Acids Symp Ser (Oxf). 2004;(48):223-4. doi: 10.1093/nass/48.1.223. Nucleic Acids Symp Ser (Oxf). 2004. PMID: 17150559
-
Investigations toward the selection of fully-modified 4'-thioRNA aptamers: optimization of in vitro transcription steps in the presence of 4'-thioNTPs.Bioorg Med Chem. 2008 Nov 1;16(21):9450-6. doi: 10.1016/j.bmc.2008.09.048. Epub 2008 Sep 20. Bioorg Med Chem. 2008. PMID: 18835173
-
Synthesis and RNA polymerase incorporation of the degenerate ribonucleotide analogue rPTP.Nucleic Acids Res. 1998 May 1;26(9):2105-11. doi: 10.1093/nar/26.9.2105. Nucleic Acids Res. 1998. PMID: 9547267 Free PMC article.
-
Regulation of bacterial growth, RNA, and protein synthesis.Annu Rev Microbiol. 1978;32:393-432. doi: 10.1146/annurev.mi.32.100178.002141. Annu Rev Microbiol. 1978. PMID: 360971 Review. No abstract available.
-
From oligonucleotide shapes to genomic SELEX: novel biological regulatory loops.Proc Natl Acad Sci U S A. 1997 Jan 7;94(1):59-64. doi: 10.1073/pnas.94.1.59. Proc Natl Acad Sci U S A. 1997. PMID: 8990161 Free PMC article. Review.
Cited by
-
Generation of ssDNA aptamers as diagnostic tool for Newcastle avian virus.PLoS One. 2020 Aug 13;15(8):e0237253. doi: 10.1371/journal.pone.0237253. eCollection 2020. PLoS One. 2020. PMID: 32790805 Free PMC article.
-
G-quadruplex induced stabilization by 2'-deoxy-2'-fluoro-D-arabinonucleic acids (2'F-ANA).Nucleic Acids Res. 2007;35(15):4977-88. doi: 10.1093/nar/gkm520. Epub 2007 Jul 17. Nucleic Acids Res. 2007. PMID: 17636049 Free PMC article.
-
In vitro selection using modified or unnatural nucleotides.Curr Protoc Nucleic Acid Chem. 2002 Feb;Chapter 9:Unit 9.6. doi: 10.1002/0471142700.nc0906s07. Curr Protoc Nucleic Acid Chem. 2002. PMID: 18428900 Review.
-
Selection, Characterization and Application of Artificial DNA Aptamer Containing Appended Bases with Sub-nanomolar Affinity for a Salivary Biomarker.Sci Rep. 2017 Mar 3;7:42716. doi: 10.1038/srep42716. Sci Rep. 2017. PMID: 28256555 Free PMC article.
-
In vitro evolution of chemically-modified nucleic acid aptamers: Pros and cons, and comprehensive selection strategies.RNA Biol. 2016 Dec;13(12):1232-1245. doi: 10.1080/15476286.2016.1236173. Epub 2016 Oct 7. RNA Biol. 2016. PMID: 27715478 Free PMC article. Review.
References
-
- Tuerk C., Gold L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science. 1990;249:505–510. - PubMed
-
- Robertson D.L., Joyce G.F. Selection in vitro of an RNA enzyme that specifically cleaves single-stranded DNA. Nature. 1990;344:467–468. - PubMed
-
- Ellington A.D., Szostak J.W. In vitro selection of RNA molecules that bind specific ligands. Nature. 1990;346:818–822. - PubMed
-
- Breaker R.R. In vitro selection of catalytic polynucleotides. Chem. Rev. 1997;97:371–390. - PubMed
-
- Stuhlmann F., Jäschke A. Characterization of an RNA active site: interactions between a Diels–Alderase ribozyme and its substrates and products. J. Am. Chem. Soc. 2002;124:3238–3244. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

