Structural bioinformatics-based design of selective, irreversible kinase inhibitors
- PMID: 15919995
- PMCID: PMC3641834
- DOI: 10.1126/science1108367
Structural bioinformatics-based design of selective, irreversible kinase inhibitors
Abstract
The active sites of 491 human protein kinase domains are highly conserved, which makes the design of selective inhibitors a formidable challenge. We used a structural bioinformatics approach to identify two selectivity filters, a threonine and a cysteine, at defined positions in the active site of p90 ribosomal protein S6 kinase (RSK). A fluoromethylketone inhibitor, designed to exploit both selectivity filters, potently and selectively inactivated RSK1 and RSK2 in mammalian cells. Kinases with only one selectivity filter were resistant to the inhibitor, yet they became sensitized after genetic introduction of the second selectivity filter. Thus, two amino acids that distinguish RSK from other protein kinases are sufficient to confer inhibitor sensitivity.
Figures


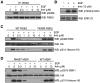
Comment in
-
Cell biology. Lessons in rational drug design for protein kinases.Science. 2005 May 27;308(5726):1266-7. doi: 10.1126/science.1113707. Science. 2005. PMID: 15919981 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Chemical Information

