Proximity of the U12 snRNA with both the 5' splice site and the branch point during early stages of spliceosome assembly
- PMID: 15923601
- PMCID: PMC1140575
- DOI: 10.1128/MCB.25.12.4813-4825.2005
Proximity of the U12 snRNA with both the 5' splice site and the branch point during early stages of spliceosome assembly
Abstract
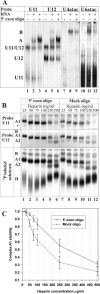
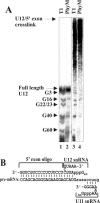
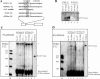
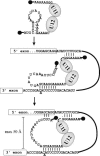
U12 snRNA is required for branch point recognition in the U12-dependent spliceosome. Using site-specific cross-linking, we have captured an unexpected interaction between the 5' end of the U12 snRNA and the -2 position upstream of the 5' splice site of P120 and SCN4a splicing substrates. The U12 snRNA nucleotides that contact the 5' exon are the same ones that form the catalytically important helix Ib with U6atac snRNA in the spliceosome catalytic core. However, the U12/5' exon interaction is transient, occurring prior to the entry of the U4atac/U6atac.U5 tri-snRNP to the spliceosome. This suggests that the helix Ib region of U12 snRNA is positioned near the 5' splice site early during spliceosome assembly and only later interacts with U6atac to form helix Ib. We also provide evidence that U12 snRNA can simultaneously interact with 5' exon sequences near 5' splice site and the branch point sequence, suggesting that the 5' splice site and branch point sequences are separated by <40 to 50 A in the complex A of the U12-dependent spliceosome. Thus, no major rearrangements are subsequently needed to position these sites for the first step of catalysis.
Figures




Similar articles
-
snRNA interactions at 5' and 3' splice sites monitored by photoactivated crosslinking in yeast spliceosomes.RNA. 1995 Nov;1(9):968-80. RNA. 1995. PMID: 8548661 Free PMC article.
-
Dynamic exchanges of RNA interactions leading to catalytic core formation in the U12-dependent spliceosome.Mol Cell. 2001 Jan;7(1):217-26. doi: 10.1016/s1097-2765(01)00169-1. Mol Cell. 2001. PMID: 11172726
-
RS domain-splicing signal interactions in splicing of U12-type and U2-type introns.Nat Struct Mol Biol. 2007 Jul;14(7):597-603. doi: 10.1038/nsmb1263. Epub 2007 Jul 1. Nat Struct Mol Biol. 2007. PMID: 17603499
-
Splicing of a rare class of introns by the U12-dependent spliceosome.Biol Chem. 2005 Aug;386(8):713-24. doi: 10.1515/BC.2005.084. Biol Chem. 2005. PMID: 16201866 Review.
-
Recognition of the 5' splice site by the spliceosome.Acta Biochim Pol. 1998;45(4):869-81. Acta Biochim Pol. 1998. PMID: 10397335 Review.
Cited by
-
Structural and mechanistic insights into human splicing factor SF3b complex derived using an integrated approach guided by the cryo-EM density maps.RNA Biol. 2016 Oct 2;13(10):1025-1040. doi: 10.1080/15476286.2016.1218590. Epub 2016 Aug 11. RNA Biol. 2016. PMID: 27618338 Free PMC article.
-
Defective minor spliceosome mRNA processing results in isolated familial growth hormone deficiency.EMBO Mol Med. 2014 Mar;6(3):299-306. doi: 10.1002/emmm.201303573. Epub 2014 Jan 30. EMBO Mol Med. 2014. PMID: 24480542 Free PMC article.
-
Tudor staphylococcal nuclease (Tudor-SN) participates in small ribonucleoprotein (snRNP) assembly via interacting with symmetrically dimethylated Sm proteins.J Biol Chem. 2012 May 25;287(22):18130-41. doi: 10.1074/jbc.M111.311852. Epub 2012 Apr 9. J Biol Chem. 2012. PMID: 22493508 Free PMC article.
-
The U1, U2 and U5 snRNAs crosslink to the 5' exon during yeast pre-mRNA splicing.Nucleic Acids Res. 2008 Feb;36(3):814-25. doi: 10.1093/nar/gkm1098. Epub 2007 Dec 15. Nucleic Acids Res. 2008. PMID: 18084028 Free PMC article.
-
The U11-48K protein contacts the 5' splice site of U12-type introns and the U11-59K protein.Mol Cell Biol. 2008 May;28(10):3548-60. doi: 10.1128/MCB.01928-07. Epub 2008 Mar 17. Mol Cell Biol. 2008. PMID: 18347052 Free PMC article.
References
-
- Abovich, N., and M. Rosbash. 1997. Cross-intron bridging interactions in the yeast commitment complex are conserved in mammals. Cell 89:403-412. - PubMed
-
- Ast, G., and A. M. Weiner. 1996. A U1/U4/U5 snRNP complex induced by a 2′-O-methyl-oligoribonucleotide complementary to U5 snRNA. Science 272:881-884. - PubMed
-
- Brow, D. A. 2002. Allosteric cascade of spliceosome activation. Annu. Rev. Genet. 36:333-360. - PubMed
-
- Burge, C. B., T. Tuschl, and P. A. Sharp. 1999. Splicing of precursors to mRNAs by the spliceosome, p. 525-560. In R. F. Gesteland, T. R. Cech, and J. F. Atkins (ed.), The RNA world, vol. 2. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
-
- Cunningham, S. A., A. J. Else, B. V. Potter, and I. C. Eperon. 1991. Influences of separation and adjacent sequences on the use of alternative 5′ splice sites. J. Mol. Biol. 217:265-281. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
