Spindle checkpoint signaling requires the mis6 kinetochore subcomplex, which interacts with mad2 and mitotic spindles
- PMID: 15930132
- PMCID: PMC1182306
- DOI: 10.1091/mbc.e05-01-0014
Spindle checkpoint signaling requires the mis6 kinetochore subcomplex, which interacts with mad2 and mitotic spindles
Abstract
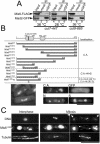
The spindle checkpoint coordinates cell cycle progression and chromosome segregation by inhibiting anaphase promoting complex/cyclosome until all kinetochores interact with the spindle properly. During early mitosis, the spindle checkpoint proteins, such as Mad2 and Bub1, accumulate at kinetochores that do not associate with the spindle. Here, we assess the requirement of various kinetochore components for the accumulation of Mad2 and Bub1 on the kinetochore in fission yeast and show that the necessity of the Mis6-complex and the Nuf2-complex is an evolutionarily conserved feature in the loading of Mad2 onto the kinetochore. Furthermore, we demonstrated that Nuf2 is required for maintaining the Mis6-complex on the kinetochore during mitosis. The Mis6-complex physically interacts with Mad2 under the condition that the Mad2-dependent checkpoint is activated. Ectopically expressed N-terminal fragments of Mis6 localize along the mitotic spindle, highlighting the potential binding ability of Mis6 not only to the centromeric chromatin but also to the spindle microtubules. We propose that the Mis6-complex, in collaboration with the Nuf2-complex, monitors the spindle-kinetochore attachment state and acts as a platform for Mad2 to accumulate at unattached kinetochores.
Figures




Similar articles
-
Dual regulation of Mad2 localization on kinetochores by Bub1 and Dam1/DASH that ensure proper spindle interaction.Mol Biol Cell. 2008 Sep;19(9):3885-97. doi: 10.1091/mbc.e08-03-0298. Epub 2008 Jul 16. Mol Biol Cell. 2008. PMID: 18632983 Free PMC article.
-
Centromere-tethered Mps1 pombe homolog (Mph1) kinase is a sufficient marker for recruitment of the spindle checkpoint protein Bub1, but not Mad1.Proc Natl Acad Sci U S A. 2012 Jan 3;109(1):209-14. doi: 10.1073/pnas.1114647109. Epub 2011 Dec 19. Proc Natl Acad Sci U S A. 2012. PMID: 22184248 Free PMC article.
-
Mad1 contribution to spindle assembly checkpoint signalling goes beyond presenting Mad2 at kinetochores.EMBO Rep. 2014 Mar;15(3):291-8. doi: 10.1002/embr.201338114. Epub 2014 Jan 29. EMBO Rep. 2014. PMID: 24477934 Free PMC article.
-
Bub1 and BubR1: at the interface between chromosome attachment and the spindle checkpoint.Mol Cell Biol. 2011 Aug;31(15):3085-93. doi: 10.1128/MCB.05326-11. Epub 2011 May 31. Mol Cell Biol. 2011. PMID: 21628528 Free PMC article. Review.
-
How the SAC gets the axe: Integrating kinetochore microtubule attachments with spindle assembly checkpoint signaling.Bioarchitecture. 2015;5(1-2):1-12. doi: 10.1080/19490992.2015.1090669. Epub 2015 Oct 2. Bioarchitecture. 2015. PMID: 26430805 Free PMC article. Review.
Cited by
-
The ABCs of CENPs.Chromosoma. 2011 Oct;120(5):425-46. doi: 10.1007/s00412-011-0330-0. Epub 2011 Jul 13. Chromosoma. 2011. PMID: 21751032 Review.
-
The kinetochore protein Kis1/Eic1/Mis19 ensures the integrity of mitotic spindles through maintenance of kinetochore factors Mis6/CENP-I and CENP-A.PLoS One. 2014 Nov 6;9(11):e111905. doi: 10.1371/journal.pone.0111905. eCollection 2014. PLoS One. 2014. PMID: 25375240 Free PMC article.
-
Dissection of CENP-C-directed centromere and kinetochore assembly.Mol Biol Cell. 2009 Oct;20(19):4246-55. doi: 10.1091/mbc.e09-05-0378. Epub 2009 Jul 29. Mol Biol Cell. 2009. PMID: 19641019 Free PMC article.
-
Biphasic incorporation of centromeric histone CENP-A in fission yeast.Mol Biol Cell. 2008 Feb;19(2):682-90. doi: 10.1091/mbc.e07-05-0504. Epub 2007 Dec 12. Mol Biol Cell. 2008. PMID: 18077559 Free PMC article.
-
Novel mad2 alleles isolated in a Schizosaccharomyces pombe gamma-tubulin mutant are defective in metaphase arrest activity, but remain functional for chromosome stability in unperturbed mitosis.Genetics. 2007 Apr;175(4):1571-84. doi: 10.1534/genetics.106.061309. Epub 2007 Feb 4. Genetics. 2007. PMID: 17277378 Free PMC article.
References
-
- Appelgren, H., Kniola, B., and Ekwall, K. (2003). Distinct centromere domain structures with separate functions demonstrated in live fission yeast cells. J. Cell Sci. 116, 4035–4042. - PubMed
-
- Cahill, D. P., Lengauer, C., Yu, J., Riggins, G. J., Willson, J. K., Markowitz, S. D., Kinzler, K. W., and Vogelstein, B. (1998). Mutations of mitotic checkpoint genes in human cancers. Nature 392, 300–303. - PubMed
-
- Cheeseman, I. M., Anderson, S., Jwa, M., Green, E. M., Kang, J., Yates, J. R., 3rd, Chan, C. S., Drubin, D. G., and Barnes, G. (2002a). Phospho-regulation of kinetochore-microtubule attachments by the Aurora kinase Ipl1p. Cell 111, 163–172. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

