oPOSSUM: identification of over-represented transcription factor binding sites in co-expressed genes
- PMID: 15933209
- PMCID: PMC1142402
- DOI: 10.1093/nar/gki624
oPOSSUM: identification of over-represented transcription factor binding sites in co-expressed genes
Abstract
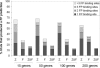
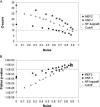
Targeted transcript profiling studies can identify sets of co-expressed genes; however, identification of the underlying functional mechanism(s) is a significant challenge. Established methods for the analysis of gene annotations, particularly those based on the Gene Ontology, can identify functional linkages between genes. Similar methods for the identification of over-represented transcription factor binding sites (TFBSs) have been successful in yeast, but extension to human genomics has largely proved ineffective. Creation of a system for the efficient identification of common regulatory mechanisms in a subset of co-expressed human genes promises to break a roadblock in functional genomics research. We have developed an integrated system that searches for evidence of co-regulation by one or more transcription factors (TFs). oPOSSUM combines a pre-computed database of conserved TFBSs in human and mouse promoters with statistical methods for identification of sites over-represented in a set of co-expressed genes. The algorithm successfully identified mediating TFs in control sets of tissue-specific genes and in sets of co-expressed genes from three transcript profiling studies. Simulation studies indicate that oPOSSUM produces few false positives using empirically defined thresholds and can tolerate up to 50% noise in a set of co-expressed genes.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

