The transcription termination factor Rho is essential and autoregulated in Caulobacter crescentus
- PMID: 15937192
- PMCID: PMC1151746
- DOI: 10.1128/JB.187.12.4290-4294.2005
The transcription termination factor Rho is essential and autoregulated in Caulobacter crescentus
Abstract
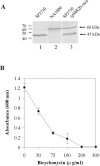
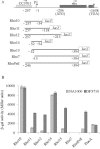
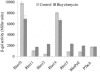
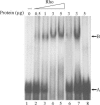
The impossibility of obtaining a rho null mutant and sensitivity to bicyclomycin have indicated that rho is essential for the viability of Caulobacter crescentus. Transcription gene fusions of sequences with serial deletions of the rho 5' untranslated region (5'-UTR) with a lacZ reporter gene indicated that rho is autoregulated at the level of attenuation of transcription in the 5'-UTR.
Figures

Similar articles
-
The conserved flaF gene has a critical role in coupling flagellin translation and assembly in Caulobacter crescentus.Mol Microbiol. 2005 Aug;57(4):1127-42. doi: 10.1111/j.1365-2958.2005.04745.x. Mol Microbiol. 2005. PMID: 16091049
-
Transcriptional regulation of the S-layer protein type I secretion system in Caulobacter crescentus.FEMS Microbiol Lett. 2005 Oct 1;251(1):29-36. doi: 10.1016/j.femsle.2005.07.028. FEMS Microbiol Lett. 2005. PMID: 16111836
-
rho is not essential for viability or virulence in Staphylococcus aureus.Antimicrob Agents Chemother. 2001 Apr;45(4):1099-103. doi: 10.1128/AAC.45.4.1099-1103.2001. Antimicrob Agents Chemother. 2001. PMID: 11257021 Free PMC article.
-
Cycle-regulated genes and cell cycle regulation.Bioessays. 2001 Jul;23(7):563-5. doi: 10.1002/bies.1081. Bioessays. 2001. PMID: 11462209 Review.
-
[Bacterial transcription termination].Postepy Biochem. 1997;43(3):182-8. Postepy Biochem. 1997. PMID: 9558707 Review. Polish. No abstract available.
Cited by
-
Temporal proteomic profiling reveals changes that support Burkholderia biofilms.Pathog Dis. 2019 Mar 1;77(2):ftz005. doi: 10.1093/femspd/ftz005. Pathog Dis. 2019. PMID: 30759239 Free PMC article.
-
Pathological R-loops in bacteria from engineered expression of endogenous antisense RNAs whose synthesis is ordinarily terminated by Rho.Nucleic Acids Res. 2024 Nov 11;52(20):12438-12455. doi: 10.1093/nar/gkae839. Nucleic Acids Res. 2024. PMID: 39373509 Free PMC article.
-
Association of the Cold Shock DEAD-Box RNA Helicase RhlE to the RNA Degradosome in Caulobacter crescentus.J Bacteriol. 2017 Jun 13;199(13):e00135-17. doi: 10.1128/JB.00135-17. Print 2017 Jul 1. J Bacteriol. 2017. PMID: 28396352 Free PMC article.
-
Mastering the control of the Rho transcription factor for biotechnological applications.Appl Microbiol Biotechnol. 2021 May;105(10):4053-4071. doi: 10.1007/s00253-021-11326-7. Epub 2021 May 8. Appl Microbiol Biotechnol. 2021. PMID: 33963893 Review.
References
-
- Alifano, P., F. Rivellini, D. Limauro, C. B. Bruni, and M. S. Carlomagno. 1991. A consensus motif common to all Rho-dependent prokaryotic transcription terminators. Cell 64:553-563. - PubMed
-
- Barik, S., P. Bhattacharya, and A. Das. 1985. Autogenous regulation of transcription termination factor Rho. J. Mol. Biol. 182:495-508. - PubMed
-
- Bektesh, S. L., and J. P. Richardson. 1980. A rho-recognition site on phage lambda cro-gene mRNA. Nature 283:102-104. - PubMed
-
- Brennan, C. A., A. J. Dombroski, and T. Platt. 1987. Transcription termination factor rho is an RNA-DNA helicase. Cell 48:945-952. - PubMed
-
- Chen, X., and B. L. Stitt. 2004. The binding of C10 oligomers to Escherichia coli transcription termination factor Rho. J. Biol. Chem. 279:16301-16310. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

