The zinc finger transcription factor Klf7 is required for TrkA gene expression and development of nociceptive sensory neurons
- PMID: 15937222
- PMCID: PMC1142558
- DOI: 10.1101/gad.1227705
The zinc finger transcription factor Klf7 is required for TrkA gene expression and development of nociceptive sensory neurons
Abstract
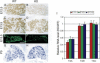
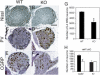
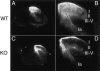
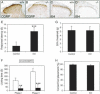
TrkA, the high affinity receptor for nerve growth factor (NGF), is essential for the development of nociceptive sensory and sympathetic neurons. The zinc finger transcription factor Klf7 interacts with an important cis element of the TrkA minimal enhancer and is coexpressed with TrkA in these neurons. We show that Klf7 binds to the endogenous TrkA minimal enhancer and can activate transcription from the TrkA minimal enhancer in a sequence-dependent manner. In Klf7(-/-) newborn mice, we find a significant reduction in sensory neurons due to increased apoptosis. The neuronal loss is restricted to nociceptive neurons that normally depend on TrkA for neurotrophic support, while other populations of somatosensory neurons appear normal. The reduction of TrkA expression in sensory neurons is a direct effect of Klf7 gene ablation, rather than a secondary effect of cell death. As a result, Klf7(-/-) mice have deficient response to noxious stimuli. Finally, removal of one TrkA allele exacerbates the loss of TrkA(+) neurons in Klf7(-/-) mice. Thus, Klf7 specifically regulates TrkA gene expression and is required for the development of a subset of nociceptive sensory neurons.
Figures




Similar articles
-
Brn3a and Klf7 cooperate to control TrkA expression in sensory neurons.Dev Biol. 2006 Dec 15;300(2):758-69. doi: 10.1016/j.ydbio.2006.08.062. Epub 2006 Aug 30. Dev Biol. 2006. PMID: 17011544
-
mKlf7, a potential transcriptional regulator of TrkA nerve growth factor receptor expression in sensory and sympathetic neurons.Development. 2001 Apr;128(7):1147-58. doi: 10.1242/dev.128.7.1147. Development. 2001. PMID: 11245580
-
Brn3a regulation of TrkA/NGF receptor expression in developing sensory neurons.Development. 2003 Aug;130(15):3525-34. doi: 10.1242/dev.00582. Development. 2003. PMID: 12810599
-
Generation of somatic sensory neuron diversity and implications on sensory coding.Curr Opin Neurobiol. 2011 Feb;21(1):52-60. doi: 10.1016/j.conb.2010.09.003. Epub 2010 Oct 1. Curr Opin Neurobiol. 2011. PMID: 20888752 Free PMC article. Review.
-
[Research progress of Krüppel-like factor 7].Sheng Li Xue Bao. 2016 Dec 25;68(6):809-815. Sheng Li Xue Bao. 2016. PMID: 28004076 Review. Chinese.
Cited by
-
Association of variants in the KIF1A gene with amyotrophic lateral sclerosis.Transl Neurodegener. 2022 Oct 26;11(1):46. doi: 10.1186/s40035-022-00320-2. Transl Neurodegener. 2022. PMID: 36284339 Free PMC article.
-
KLF7/VPS35 axis contributes to hepatocellular carcinoma progression through CCDC85C-activated β-catenin pathway.Cell Biosci. 2021 Apr 15;11(1):73. doi: 10.1186/s13578-021-00585-6. Cell Biosci. 2021. PMID: 33858520 Free PMC article.
-
Signals and switches in Mammalian neural crest cell differentiation.Cold Spring Harb Perspect Biol. 2013 Feb 1;5(2):a008326. doi: 10.1101/cshperspect.a008326. Cold Spring Harb Perspect Biol. 2013. PMID: 23378583 Free PMC article. Review.
-
KLF7 promotes pancreatic cancer growth and metastasis by up-regulating ISG expression and maintaining Golgi complex integrity.Proc Natl Acad Sci U S A. 2020 Jun 2;117(22):12341-12351. doi: 10.1073/pnas.2005156117. Epub 2020 May 19. Proc Natl Acad Sci U S A. 2020. PMID: 32430335 Free PMC article.
-
Krüppel-like factor 6 rendered rat Schwann cell more sensitive to apoptosis via upregulating FAS expression.PLoS One. 2013 Dec 4;8(12):e82449. doi: 10.1371/journal.pone.0082449. eCollection 2013. PLoS One. 2013. PMID: 24324791 Free PMC article.
References
-
- Alberti L., Carniti, C., Miranda, C., Roccato, E., and Pierotti, M.A. 2003. RET and NTRK1 proto-oncogenes in human diseases. J. Cell Physiol. 195: 168-186. - PubMed
-
- Anderson D.J. 1997. Cellular and molecular biology of neural crest cell lineage determination. Trends Genet. 13: 276-280. - PubMed
-
- Bibel M. and Barde, Y.A. 2000. Neurotrophins: Key regulators of cell fate and cell shape in the vertebrate nervous system. Genes & Dev. 14: 2919-2937. - PubMed
-
- Black A.R., Black, J.D., and Azizkhan-Clifford, J. 2001. Sp1 and kruppel-like factor family of transcription factors in cell growth regulation and cancer. J. Cell Physiol. 188: 143-160. - PubMed
-
- Chen Z.F., Rebelo, S., White, F., Malmberg, A.B., Baba, H., Lima, D., Woolf, C.J., Basbaum, A.I., and Anderson, D.J. 2001. The paired homeodomain protein DRG11 is required for the projection of cutaneous sensory afferent fibers to the dorsal spinal cord. Neuron 31: 59-73. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
