DNA damage induced hyperphosphorylation of replication protein A. 1. Identification of novel sites of phosphorylation in response to DNA damage
- PMID: 15938632
- PMCID: PMC4331072
- DOI: 10.1021/bi0480584
DNA damage induced hyperphosphorylation of replication protein A. 1. Identification of novel sites of phosphorylation in response to DNA damage
Abstract
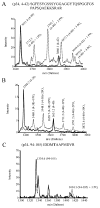
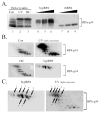
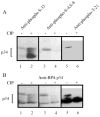
Replication protein A (RPA) is the predominant eukaryotic single-stranded DNA binding protein composed of 70, 34, and 14 kDa subunits. RPA plays central roles in the processes of DNA replication, repair, and recombination, and the p34 subunit of RPA is phosphorylated in a cell-cycle-dependent fashion and is hyperphosphorylated in response to DNA damage. We have developed an in vitro procedure for the preparation of hyperphosphorylated RPA and characterized a series of novel sites of phosphorylation using a combination of in gel tryptic digestion, SDS-PAGE and HPLC, MALDI-TOF MS analysis, 2D gel electrophoresis, and phosphospecific antibodies. We have mapped five phosphorylation sites on the RPA p34 subunit and five sites of phosphorylation on the RPA p70 subunit. No modification of the 14 kDa subunit was observed. Using the procedures developed with in vitro phosphorylated RPA, we confirmed a series of phosphorylation events on RPA from HeLa cells that was hyperphosphorylated in vivo in response to the DNA damaging agents, aphidicolin and hydroxyurea.
Figures




References
-
- Braun K, Lao Y, He Z, Ingles C, Wold M. Role of protein–protein interactions in the function of replication protein A (RPA): RPA modulates the activity of DNA polymerase alpha by multiple mechanisms. Biochemistry. 1997;36:8443–8454. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

