Functional analysis of the N- and C-terminus of mammalian G9a histone H3 methyltransferase
- PMID: 15939934
- PMCID: PMC1143580
- DOI: 10.1093/nar/gki635
Functional analysis of the N- and C-terminus of mammalian G9a histone H3 methyltransferase
Abstract
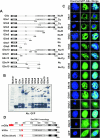
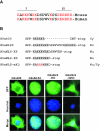
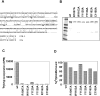
Methylation of lysine 9 (K9) in the N-terminus tail of histone H3 (H3) in chromatin is associated with transcriptionally silenced genes and is mediated by histone methyltransferases. Murine G9a is a 1263 amino acid H3-K9 methyltransferase that possesses characteristic SET domain and ANK repeats. In this paper, we have used a series of green fluorescent protein-tagged deletion constructs to identify two nuclear localization signals (NLS), the first NLS embedded between amino acids 24 and 109 and the second between amino acids 394 and 401 of murine G9a. Our data show that both long and short G9a isoforms were capable of entering the nucleus to methylate chromatin. Full-length or N-terminus-deleted G9a isoforms were also catalytically active enzymes that methylated recombinant H3 or synthetic peptides representing the N-terminus tail of H3. In vitro methylation reactions using N-terminus tail peptides resulted in tri-methylation of K9 that remained processive, even in G9a enzymes that lacked an N-terminus region by deletion. Co-expression of G9a and H3 resulted in di- and tri-methylation of H3-K9, while siRNA-mediated knockdown of G9a in HeLa cells resulted in reduction of global H3-K9 di- and tri-methylation. A recombinant deletion mutant enzyme fused with maltose-binding protein (MBP-G9aDelta634) was used for steady-state kinetic analysis with various substrates and was compared with full-length G9a (G9aFL). Turnover numbers of MBP-G9aDelta634 for various substrates was approximately 3-fold less compared with G9aFL, while their Michaelis constants (K(m)) for recombinant H3 were similar. The K(AdoMet)m for MBP-G9aDelta634 was approximately 2.3-2.65 microM with various substrates. Catalytic efficiencies (kcat/K(m)) for both MBP-G9aDelta634 and G9aFL were similar, suggesting that the N-terminus is not essential for catalysis. Furthermore, mutation of conserved amino acids R1097A, W1103A, Y1120A, Y1138A and R1162A, or the metal binding C1168A in the catalytic region, resulted in catalytically impaired enzymes, thereby confirming the involvement of the C-terminus of G9a in catalysis. Thus, distinct domains modulate nuclear targeting and catalytic functions of G9a.
Figures


Similar articles
-
Sequence specificity and role of proximal amino acids of the histone H3 tail on catalysis of murine G9A lysine 9 histone H3 methyltransferase.Biochemistry. 2005 Oct 4;44(39):12998-3006. doi: 10.1021/bi0509907. Biochemistry. 2005. PMID: 16185068
-
Catalytic properties and kinetic mechanism of human recombinant Lys-9 histone H3 methyltransferase SUV39H1: participation of the chromodomain in enzymatic catalysis.Biochemistry. 2006 Mar 14;45(10):3272-84. doi: 10.1021/bi051997r. Biochemistry. 2006. PMID: 16519522
-
Substrate specificity and kinetic mechanism of mammalian G9a histone H3 methyltransferase.J Biol Chem. 2004 Dec 17;279(51):53248-58. doi: 10.1074/jbc.M409604200. Epub 2004 Oct 14. J Biol Chem. 2004. PMID: 15485804
-
Structure and function of histone H3 lysine 9 methyltransferases and demethylases.Chembiochem. 2011 Jan 24;12(2):254-63. doi: 10.1002/cbic.201000545. Epub 2011 Jan 5. Chembiochem. 2011. PMID: 21243713 Review.
-
The many faces of histone lysine methylation.Curr Opin Cell Biol. 2002 Jun;14(3):286-98. doi: 10.1016/s0955-0674(02)00335-6. Curr Opin Cell Biol. 2002. PMID: 12067650 Review.
Cited by
-
G9a, a multipotent regulator of gene expression.Epigenetics. 2013 Jan;8(1):16-22. doi: 10.4161/epi.23331. Epub 2012 Dec 20. Epigenetics. 2013. PMID: 23257913 Free PMC article. Review.
-
UHRF1 binds G9a and participates in p21 transcriptional regulation in mammalian cells.Nucleic Acids Res. 2009 Feb;37(2):493-505. doi: 10.1093/nar/gkn961. Epub 2008 Dec 4. Nucleic Acids Res. 2009. PMID: 19056828 Free PMC article.
-
Lysine methyltransferase G9a is not required for DNMT3A/3B anchoring to methylated nucleosomes and maintenance of DNA methylation in somatic cells.Epigenetics Chromatin. 2012 Jan 27;5(1):3. doi: 10.1186/1756-8935-5-3. Epigenetics Chromatin. 2012. PMID: 22284370 Free PMC article.
-
Identification of three histone methyltransferases in Drosophila: dG9a is a suppressor of PEV and is required for gene silencing.Mol Genet Genomics. 2006 Jun;275(6):513-26. doi: 10.1007/s00438-006-0116-x. Epub 2006 Apr 19. Mol Genet Genomics. 2006. PMID: 16622709
-
Transcriptional repressor REST drives lineage stage-specific chromatin compaction at Ptch1 and increases AKT activation in a mouse model of medulloblastoma.Sci Signal. 2019 Jan 22;12(565):eaan8680. doi: 10.1126/scisignal.aan8680. Sci Signal. 2019. PMID: 30670636 Free PMC article.
References
-
- Turner B.M. Histone acetylation and an epigenetic code. Bioessays. 2000;22:836–845. - PubMed
-
- Karpen G.H., Le M.H., Le H. Centric heterochromatin and the efficiency of achiasmate disjunction in Drosophila female meiosis. Science. 1996;273:118–122. - PubMed
-
- Dernburg A.F., Sedat J.W., Hawley R.S. Direct evidence of a role for heterochromatin in meiotic chromosome segregation. Cell. 1996;86:135–146. - PubMed
-
- Moazed D., Rudner A.D., Huang J., Hoppe G.J., Tanny J.C. A model for step-wise assembly of heterochromatin in yeast. Novartis Found Symp. 2004;259:48–56. discussion 56–62, 163–169. - PubMed
-
- Okuwaki M., Verreault A. Maintenance DNA methylation of nucleosome core particles. J. Biol. Chem. 2004;279:2904–2912. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

