DNA sequence polymorphism and divergence at the erect wing and suppressor of sable loci of Drosophila melanogaster and D. simulans
- PMID: 15944367
- PMCID: PMC1451169
- DOI: 10.1534/genetics.104.033456
DNA sequence polymorphism and divergence at the erect wing and suppressor of sable loci of Drosophila melanogaster and D. simulans
Abstract
Several evolutionary models of linked selection (e.g., genetic hitchhiking, background selection, and random environment) predict a reduction in polymorphism relative to divergence in genomic regions where the rate of crossing over per physical distance is restricted. We tested this prediction near the telomere of the Drosophila melanogaster and D. simulans X chromosome at two loci, erect wing (ewg) and suppressor of sable [su(s)]. Consistent with this prediction, polymorphism is reduced at both loci, while divergence is normal. The reduction is greater at ewg, the more distal of the two regions. Two models can be discriminated by comparing the observed site frequency spectra with those predicted by the models. The hitchhiking model predicts a skew toward rare variants in a sample, while the spectra under the background-selection model are similar to those of the neutral model of molecular evolution. Statistical tests of the fit to the predictions of these models require many sampled alleles and segregating sites. Thus we used SSCP and stratified DNA sequencing to cover a large number of randomly sampled alleles (approximately 50) from each of three populations. The result is a clear trend toward negative values of Tajima's D, indicating an excess of rare variants at ewg, the more distal of the two loci. One fixed difference among the populations and high FST values indicate strong population subdivision among the three populations at ewg. These results indicate genetic hitchhiking at ewg, in particular, geographically localized hitchhiking events within Africa. The reduction of polymorphism at su(s) combined with the excess of high-frequency variants in D. simulans is inconsistent with the hitchhiking and background-selection models.
Figures
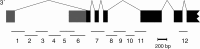

References
-
- Aguadé, M., W. Meyers, A. D. Long and C. H. Langley, 1994. Single-strand conformation polymorphism analysis coupled with stratified DNA sequencing reveals reduced sequence variation in the su(s) and su(wa) regions of the Drosophila melanogaster X chromosome. Proc. Natl. Acad. Sci. USA 91 4658–4662. - PMC - PubMed
-
- Andolfatto, P., 2001. Contrasting patterns of X-linked and autosomal nucleotide variation in Drosophila melanogaster and Drosophila simulans. Mol. Biol. Evol. 18 279–290. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

