Different cytokine response of primary colonic epithelial cells to commensal bacteria
- PMID: 15948242
- PMCID: PMC4315991
- DOI: 10.3748/wjg.v11.i22.3375
Different cytokine response of primary colonic epithelial cells to commensal bacteria
Abstract
Aim: To determine if primary murine colonic epithelial cells (CEC) respond to commensal bacteria and discriminate between different types of bacteria.
Methods: A novel CEC: bacteria co-culture system was used to compare the ability of the colonic commensal bacteria, Bacteroides ovatus, E. coli (SLF) and Lactobacillus rhamnosus (LGG) to modulate production of different cytokines (n = 15) by primary CEC. Antibody staining and flow cytometry were used to investigate Toll-like receptor (TLR) expression by CEC directly ex vivo and TLR responsiveness was determined by examining the ability of TLR ligands to influence CEC cytokine production.
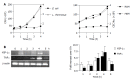
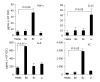
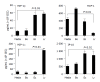
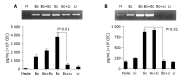
Results: Primary CEC constitutively expressed functional TLR2 and TLR4. Cultured in complete medium alone, CEC secreted IL-6, MCP-1 and IP-10 the levels of which were significantly increased upon addition of the TLR ligands peptidoglycan (PGN) and lipopolysaccharide (LPS). Exposure to the commensal bacteria induced or up-regulated different patterns of cytokine production and secretion. E. coli induced production of MIP-1alpha/beta and betadefensin3 whereas B. ovatus and L. rhamnosus exclusively induced MCP-1 and MIP-2alpha expression, respectively. TNFalpha, RANTES and MEC were induced or up-regulated in response to some but not all of the bacteria whereas ENA78 and IP-10 were up-regulated in response to all bacteria. Evidence of bacterial interference and suppression of cytokine production was obtained from mixed bacterial: CEC co-cultures. Probiotic LGG suppressed E. coli- and B. ovatus-induced cytokine mRNA accumulation and protein secretion.
Conclusion: These observations demonstrate the ability of primary CEC to respond to and discriminate between different strains of commensal bacteria and identify a mechanism by which probiotic bacteria (LGG) may exert anti-inflammatory effects in vivo.
Figures


Similar articles
-
Toll-like receptor-mediated responses of primary intestinal epithelial cells during the development of colitis.Am J Physiol Gastrointest Liver Physiol. 2005 Mar;288(3):G514-24. doi: 10.1152/ajpgi.00377.2004. Epub 2004 Oct 21. Am J Physiol Gastrointest Liver Physiol. 2005. PMID: 15499080
-
Serotonin disturbs colon epithelial tolerance of commensal E. coli by increasing NOX2-derived superoxide.Free Radic Biol Med. 2017 May;106:196-207. doi: 10.1016/j.freeradbiomed.2017.02.034. Epub 2017 Feb 17. Free Radic Biol Med. 2017. PMID: 28216386
-
Exposure of intestinal epithelial cells to UV-killed Lactobacillus GG but not Bifidobacterium breve enhances the effector immune response in vitro.Int Arch Allergy Immunol. 2010;152(2):159-68. doi: 10.1159/000265537. Epub 2009 Dec 16. Int Arch Allergy Immunol. 2010. PMID: 20016198
-
Lactobacillus rhamnosus protects human colonic muscle from pathogen lipopolysaccharide-induced damage.Neurogastroenterol Motil. 2013 Dec;25(12):984-e777. doi: 10.1111/nmo.12232. Epub 2013 Oct 1. Neurogastroenterol Motil. 2013. PMID: 24118564
-
Commensal bacteria (normal microflora), mucosal immunity and chronic inflammatory and autoimmune diseases.Immunol Lett. 2004 May 15;93(2-3):97-108. doi: 10.1016/j.imlet.2004.02.005. Immunol Lett. 2004. PMID: 15158604 Review.
Cited by
-
Lactobacillus acidophilus induces cytokine and chemokine production via NF-κB and p38 mitogen-activated protein kinase signaling pathways in intestinal epithelial cells.Clin Vaccine Immunol. 2012 Apr;19(4):603-8. doi: 10.1128/CVI.05617-11. Epub 2012 Feb 22. Clin Vaccine Immunol. 2012. PMID: 22357649 Free PMC article.
-
Lactobacillus acidophilus induces a slow but more sustained chemokine and cytokine response in naïve foetal enterocytes compared to commensal Escherichia coli.BMC Immunol. 2010 Jan 19;11:2. doi: 10.1186/1471-2172-11-2. BMC Immunol. 2010. PMID: 20085657 Free PMC article.
-
Cytokine response after stimulation of culture cells by zinc and probiotic strain.In Vitro Cell Dev Biol Anim. 2019 Dec;55(10):830-837. doi: 10.1007/s11626-019-00401-z. Epub 2019 Sep 13. In Vitro Cell Dev Biol Anim. 2019. PMID: 31520371
-
Experimental Evidence for Adaptation to Species-Specific Gut Microbiota in House Mice.mSphere. 2019 Jul 10;4(4):e00387-19. doi: 10.1128/mSphere.00387-19. mSphere. 2019. PMID: 31292233 Free PMC article.
-
Communicable ulcerative colitis induced by T-bet deficiency in the innate immune system.Cell. 2007 Oct 5;131(1):33-45. doi: 10.1016/j.cell.2007.08.017. Cell. 2007. PMID: 17923086 Free PMC article.
References
-
- Baumgart DC, Dignass AU. Intestinal barrier function. Curr Opin Clin Nutr Metab Care. 2002;5:685–694. - PubMed
-
- Hecht G. Microbes and microbial toxins: paradigms for microbial-mucosal interactions. VII. Enteropathogenic Escherichia coli: physiological alterations from an extracellular position. Am J Physiol Gastrointest Liver Physiol. 2001;281:G1–G7. - PubMed
-
- Melmed G, Thomas LS, Lee N, Tesfay SY, Lukasek K, Michelsen KS, Zhou Y, Hu B, Arditi M, Abreu MT. Human intestinal epithelial cells are broadly unresponsive to Toll-like receptor 2-dependent bacterial ligands: implications for host-microbial interactions in the gut. J Immunol. 2003;170:1406–1415. - PubMed
-
- Fusunyan RD, Nanthakumar NN, Baldeon ME, Walker WA. Evidence for an innate immune response in the immature human intestine: toll-like receptors on fetal enterocytes. Pediatr Res. 2001;49:589–593. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

