Unzipped and loaded: the role of DNA helicases and RFC clamp-loading complexes in sister chromatid cohesion
- PMID: 15955849
- PMCID: PMC2171654
- DOI: 10.1083/jcb.200503129
Unzipped and loaded: the role of DNA helicases and RFC clamp-loading complexes in sister chromatid cohesion
Abstract
It is well known that the products of chromosome replication are paired to ensure that the sisters segregate away from each other during mitosis. A key issue is how cells pair sister chromatids but preclude the catastrophic pairing of nonsister chromatids. The identification of both replication factor C and DNA helicases as critical for sister chromatid pairing has brought new insights into this fundamental process.
Figures
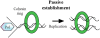
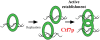
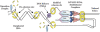
Similar articles
-
Fork it over: the cohesion establishment factor Ctf7p and DNA replication.J Cell Sci. 2007 Aug 1;120(Pt 15):2471-7. doi: 10.1242/jcs.011999. J Cell Sci. 2007. PMID: 17646671 Review.
-
Cohesin SMC1 beta is required for meiotic chromosome dynamics, sister chromatid cohesion and DNA recombination.Nat Cell Biol. 2004 Jun;6(6):555-62. doi: 10.1038/ncb1135. Epub 2004 May 16. Nat Cell Biol. 2004. PMID: 15146193
-
The mechanism of sister chromatid cohesion.Exp Cell Res. 2004 May 15;296(1):80-5. doi: 10.1016/j.yexcr.2004.03.005. Exp Cell Res. 2004. PMID: 15120997 Review.
-
Essential roles for cohesin in kinetochore and spindle function in Xenopus egg extracts.J Cell Sci. 2006 Dec 15;119(Pt 24):5057-66. doi: 10.1242/jcs.03277. J Cell Sci. 2006. PMID: 17158911
-
Establishment of sister chromatid cohesion at the S. cerevisiae replication fork.Mol Cell. 2006 Sep 15;23(6):787-99. doi: 10.1016/j.molcel.2006.08.018. Epub 2006 Sep 7. Mol Cell. 2006. PMID: 16962805
Cited by
-
Nucleolar protein trafficking in response to HIV-1 Tat: rewiring the nucleolus.PLoS One. 2012;7(11):e48702. doi: 10.1371/journal.pone.0048702. Epub 2012 Nov 15. PLoS One. 2012. PMID: 23166591 Free PMC article.
-
RFCCtf18 and the Swi1-Swi3 complex function in separate and redundant pathways required for the stabilization of replication forks to facilitate sister chromatid cohesion in Schizosaccharomyces pombe.Mol Biol Cell. 2008 Feb;19(2):595-607. doi: 10.1091/mbc.e07-06-0618. Epub 2007 Nov 28. Mol Biol Cell. 2008. PMID: 18045993 Free PMC article.
-
Roles of the sister chromatid cohesion apparatus in gene expression, development, and human syndromes.Chromosoma. 2007 Feb;116(1):1-13. doi: 10.1007/s00412-006-0072-6. Epub 2006 Jul 4. Chromosoma. 2007. PMID: 16819604 Free PMC article. Review.
-
Interaction of Deubiquitinase 2A-DUB/MYSM1 with DNA Repair and Replication Factors.Int J Mol Sci. 2020 May 26;21(11):3762. doi: 10.3390/ijms21113762. Int J Mol Sci. 2020. PMID: 32466590 Free PMC article.
-
The chromosome cycle: coordinating replication and segregation. Second in the cycles review series.EMBO Rep. 2005 Nov;6(11):1028-34. doi: 10.1038/sj.embor.7400557. EMBO Rep. 2005. PMID: 16264427 Free PMC article. Review.
References
-
- Ahkmedov, A.T., C. Frei, M. Tasi-Pflugfelder, B. Kemper, S.M. Gasser, and R. Jessberger. 1998. Structural maintenance of chromosome protein C-terminal domains bind preferentially to DNA with secondary structure. J. Biol. Chem. 273:24088–24094. - PubMed
-
- Arumugam, P., S. Gruger, K. Tanaka, C.H. Haering, K. Mechtler, and K. Nasmyth. 2003. ATP hydrolysis is required for cohesin's association with chromosomes. Curr. Biol. 13:1941–1953. - PubMed

