Junctional adhesion molecule a serves as a receptor for prototype and field-isolate strains of mammalian reovirus
- PMID: 15956543
- PMCID: PMC1143703
- DOI: 10.1128/JVI.79.13.7967-7978.2005
Junctional adhesion molecule a serves as a receptor for prototype and field-isolate strains of mammalian reovirus
Abstract
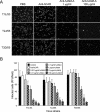
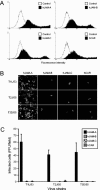
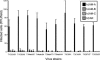
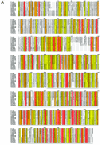
Reovirus infections are initiated by the binding of viral attachment protein sigma1 to receptors on the surface of host cells. The sigma1 protein is an elongated fiber comprised of an N-terminal tail that inserts into the virion and a C-terminal head that extends from the virion surface. The prototype reovirus strains type 1 Lang/53 (T1L/53) and type 3 Dearing/55 (T3D/55) use junctional adhesion molecule A (JAM-A) as a receptor. The C-terminal half of the T3D/55 sigma1 protein interacts directly with JAM-A, but the determinants of receptor-binding specificity have not been identified. In this study, we investigated whether JAM-A also mediates the attachment of the prototype reovirus strain type 2 Jones/55 (T2J/55) and a panel of field-isolate strains representing each of the three serotypes. Antibodies specific for JAM-A were capable of inhibiting infections of HeLa cells by T1L/53, T2J/55, and T3D/55, demonstrating that strains of all three serotypes use JAM-A as a receptor. To corroborate these findings, we introduced JAM-A or the structurally related JAM family members JAM-B and JAM-C into Chinese hamster ovary cells, which are poorly permissive for reovirus infection. Both prototype and field-isolate reovirus strains were capable of infecting cells transfected with JAM-A but not those transfected with JAM-B or JAM-C. A sequence analysis of the sigma1-encoding S1 gene segment of the strains chosen for study revealed little conservation in the deduced sigma1 amino acid sequences among the three serotypes. This contrasts markedly with the observed sequence variability within each serotype, which is confined to a small number of amino acids. Mapping of these residues onto the crystal structure of sigma1 identified regions of conservation and variability, suggesting a likely mode of JAM-A binding via a conserved surface at the base of the sigma1 head domain.
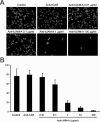
Figures



Similar articles
-
Reovirus binding determinants in junctional adhesion molecule-A.J Biol Chem. 2007 Jun 15;282(24):17930-40. doi: 10.1074/jbc.M702180200. Epub 2007 Apr 23. J Biol Chem. 2007. PMID: 17452315
-
JAM-A-independent, antibody-mediated uptake of reovirus into cells leads to apoptosis.J Virol. 2006 Feb;80(3):1261-70. doi: 10.1128/JVI.80.3.1261-1270.2006. J Virol. 2006. PMID: 16415003 Free PMC article.
-
Reovirus μ1 Protein Affects Infectivity by Altering Virus-Receptor Interactions.J Virol. 2016 Nov 14;90(23):10951-10962. doi: 10.1128/JVI.01843-16. Print 2016 Dec 1. J Virol. 2016. PMID: 27681135 Free PMC article.
-
Reovirus receptors, cell entry, and proapoptotic signaling.Adv Exp Med Biol. 2013;790:42-71. doi: 10.1007/978-1-4614-7651-1_3. Adv Exp Med Biol. 2013. PMID: 23884585 Free PMC article. Review.
-
From touchdown to transcription: the reovirus cell entry pathway.Curr Top Microbiol Immunol. 2010;343:91-119. doi: 10.1007/82_2010_32. Curr Top Microbiol Immunol. 2010. PMID: 20397070 Free PMC article. Review.
Cited by
-
Genetic determinants of reovirus pathogenesis in a murine model of respiratory infection.J Virol. 2013 Aug;87(16):9279-89. doi: 10.1128/JVI.00182-13. Epub 2013 Jun 12. J Virol. 2013. PMID: 23760238 Free PMC article.
-
Utilization of sialylated glycans as coreceptors enhances the neurovirulence of serotype 3 reovirus.J Virol. 2012 Dec;86(24):13164-73. doi: 10.1128/JVI.01822-12. Epub 2012 Oct 3. J Virol. 2012. PMID: 23035227 Free PMC article.
-
p38 Mitogen-Activated Protein Kinase Signaling Enhances Reovirus Replication by Facilitating Efficient Virus Entry, Capsid Uncoating, and Postuncoating Steps.J Virol. 2023 Feb 28;97(2):e0000923. doi: 10.1128/jvi.00009-23. Epub 2023 Feb 6. J Virol. 2023. PMID: 36744961 Free PMC article.
-
Virus-Receptor Interactions: The Key to Cellular Invasion.J Mol Biol. 2018 Aug 17;430(17):2590-2611. doi: 10.1016/j.jmb.2018.06.024. Epub 2018 Jun 18. J Mol Biol. 2018. PMID: 29924965 Free PMC article. Review.
-
Sialic Acid Receptors: The Key to Solving the Enigma of Zoonotic Virus Spillover.Viruses. 2021 Feb 8;13(2):262. doi: 10.3390/v13020262. Viruses. 2021. PMID: 33567791 Free PMC article. Review.
References
-
- Banerjea, A. C., K. A. Brechling, C. A. Ray, H. Erikson, D. J. Pickup, and W. K. Joklik. 1988. High-level synthesis of biologically active reovirus protein sigma 1 in a mammalian expression vector system. Virology 167:601-612. - PubMed
-
- Barton, E. S., J. L. Connolly, J. C. Forrest, J. D. Chappell, and T. S. Dermody. 2001. Utilization of sialic acid as a coreceptor enhances reovirus attachment by multistep adhesion strengthening. J. Biol. Chem. 276:2200-2211. - PubMed
-
- Barton, E. S., J. C. Forrest, J. L. Connolly, J. D. Chappell, Y. Liu, F. Schnell, A. Nusrat, C. A. Parkos, and T. S. Dermody. 2001. Junction adhesion molecule is a receptor for reovirus. Cell 104:441-451. - PubMed
-
- Barton, G. J. 1993. ALSCRIPT: a tool to format multiple sequence alignments. Protein Eng. 6:37-40. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

