High functional overlap between MluI cell-cycle box binding factor and Swi4/6 cell-cycle box binding factor in the G1/S transcriptional program in Saccharomyces cerevisiae
- PMID: 15965243
- PMCID: PMC1456534
- DOI: 10.1534/genetics.105.044560
High functional overlap between MluI cell-cycle box binding factor and Swi4/6 cell-cycle box binding factor in the G1/S transcriptional program in Saccharomyces cerevisiae
Abstract
In budding yeast, many genes are induced early in the cell cycle. Induction of these genes has been predominantly attributed to two transcription factors, Swi4-Swi6 (SBF) and Mbp1-Swi6 (MBF). Swi4 and Mbp1 are related DNA-binding proteins with dissimilar target sequences. For most G1/S-regulated genes that we tested in a cdc20 block-release protocol for cell-cycle synchronization, removal of both Swi4 and Mbp1 was necessary and sufficient to essentially eliminate cell-cycle-regulated expression. Detectable SBF or MBF binding sites (SCBs or MCBs) in the promoters or available genome-wide promoter occupancy data do not consistently explain this functional overlap. The overlapping ability of these transcription factors to regulate many promoters with very similar cell-cycle kinetics may provide robustness to the G1/S transcriptional response, but poses a puzzle with respect to promoter-transcription factor specificity. In addition, for some genes, deletion of Mbp1 or Swi4 enhances transcription, suggesting that these factors can also function as transcriptional repressors. Finally, we observe residual G1/S transcriptional regulation in the absence of Swi4 and Mbp1.
Figures


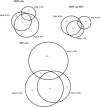

References
-
- Amon, A., M. Tyers, B. Futcher and K. Nasmyth, 1993. Mechanisms that help the yeast cell cycle clock tick: G2 cyclins transcriptionally activate G2 cyclins and repress G1 cyclins. Cell 74: 993–1007. - PubMed
-
- Andrews, B. J., and I. Herskowitz, 1989. a Identification of a DNA binding factor involved in cell-cycle control of the yeast HO gene. Cell 57: 21–29. - PubMed
-
- Andrews, B. J., and I. Herskowitz, 1989. b The yeast SWI4 protein contains a motif present in developmental regulators and is part of a complex involved in cell-cycle-dependent transcription. Nature 342: 830–833. - PubMed
-
- Breeden, L., and K. Nasmyth, 1987. Cell cycle control of the yeast HO gene: cis- and trans-acting regulators. Cell 48: 389–397. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

