Zn2+-dependent deoxyribozymes that form natural and unnatural RNA linkages
- PMID: 15966746
- PMCID: PMC1586068
- DOI: 10.1021/bi050146g
Zn2+-dependent deoxyribozymes that form natural and unnatural RNA linkages
Abstract
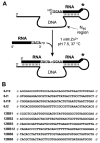
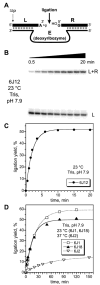
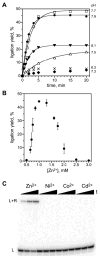
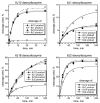
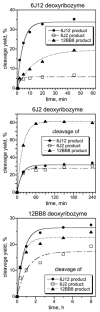
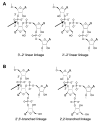
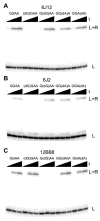
We report Zn(2+)-dependent deoxyribozymes that ligate RNA. The DNA enzymes were identified by in vitro selection and ligate RNA with k(obs) up to 0.5 min(-)(1) at 1 mM Zn(2+) and 23 degrees C, pH 7.9, which is substantially faster than our previously reported Mg(2+)-dependent deoxyribozymes. Each new Zn(2+)-dependent deoxyribozyme mediates the reaction of a specific nucleophile on one RNA substrate with a 2',3'-cyclic phosphate on a second RNA substrate. Some of the Zn(2+)-dependent deoxyribozymes create native 3'-5' RNA linkages (with k(obs) up to 0.02 min(-)(1)), whereas all of our previous Mg(2+)-dependent deoxyribozymes that use a 2',3'-cyclic phosphate create non-native 2'-5' RNA linkages. On this basis, Zn(2+)-dependent deoxyribozymes have promise for synthesis of native 3'-5'-linked RNA using 2',3'-cyclic phosphate RNA substrates, although these particular Zn(2+)-dependent deoxyribozymes are likely not useful for this practical application. Some of the new Zn(2+)-dependent deoxyribozymes instead create non-native 2'-5' linkages, just like their Mg(2+) counterparts. Unexpectedly, other Zn(2+)-dependent deoxyribozymes synthesize one of three unnatural linkages that are formed upon the reaction of an RNA nucleophile other than a 5'-hydroxyl group. Two of these unnatural linkages are the 3'-2' and 2'-2' linear junctions created when the 2'-hydroxyl of the 5'-terminal guanosine of one RNA substrate attacks the 2',3'-cyclic phosphate of the second RNA substrate. The third unnatural linkage is a branched RNA that results from attack of a specific internal 2'-hydroxyl of one RNA substrate at the 2',3'-cyclic phosphate. When compared with the consistent creation of 2'-5' linkages by Mg(2+)-dependent ligation, formation of this variety of RNA ligation products by Zn(2+)-dependent deoxyribozymes highlights the versatility of transition metals such as Zn(2+) for mediating nucleic acid catalysis.
Figures
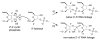




References
-
- Kruger K, Grabowski PJ, Zaug AJ, Sands J, Gottschling DE, Cech TR. Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell. 1982;31:147–157. - PubMed
-
- Cech TR. Self-splicing of group I introns. Annu Rev Biochem. 1990;59:543–568. - PubMed
-
- Qin PZ, Pyle AM. The architectural organization and mechanistic function of group II intron structural elements. Curr Opin Struct Biol. 1998;8:301–308. - PubMed
-
- Guerrier-Takada C, Gardiner K, Marsh T, Pace N, Altman S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell. 1983;35:849–857. - PubMed
-
- Breaker RR. DNA enzymes. Nature Biotechnol. 1997;15:427–431. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

