H++: a server for estimating pKas and adding missing hydrogens to macromolecules
- PMID: 15980491
- PMCID: PMC1160225
- DOI: 10.1093/nar/gki464
H++: a server for estimating pKas and adding missing hydrogens to macromolecules
Abstract
The structure and function of macromolecules depend critically on the ionization (protonation) states of their acidic and basic groups. A number of existing practical methods predict protonation equilibrium pK constants of macromolecules based upon their atomic resolution Protein Data Bank (PDB) structures; the calculations are often performed within the framework of the continuum electrostatics model. Unfortunately, these methodologies are complex, involve multiple steps and require considerable investment of effort. Our web server http://biophysics.cs.vt.edu/H++ provides access to a tool that automates this process, allowing both experts and novices to quickly obtain estimates of pKs as well as other related characteristics of biomolecules such as isoelectric points, titration curves and energies of protonation microstates. Protons are added to the input structure according to the calculated ionization states of its titratable groups at the user-specified pH; the output is in the PQR (PDB + charges + radii) format. In addition, corresponding coordinate and topology files are generated in the format supported by the molecular modeling package AMBER. The server is intended for a broad community of biochemists, molecular modelers, structural biologists and drug designers; it can also be used as an educational tool in biochemistry courses.
Figures
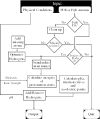
References
-
- Perutz M. Electrostatic effects in proteins. Science. 1978;201:1187–1191. - PubMed
-
- Honig B., Nicholls A. Classical electrostatics in biology and chemistry. Science. 1995;268:1144. - PubMed
-
- Davis M.E., McCammon J.A. Electrostatics in biomolecular structure and dynamics. Chem. Rev. 1990;94:7684–7692.
-
- Baker N.A., McCammon J.A. Electrostatic Interactions. In: Bourne P., Weissig H., editors. Structural Bioinformatics. NY: John Wiley & Sons, Inc.; 2002. pp. 427–440.
-
- Warshel A., Åqvist J. Electrostatic energy and macromolecular function. Ann. Rev. Biophys. Biophys. Chem. 1991;20:267–298. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

