New insights into metabolic properties of marine bacteria encoding proteorhodopsins
- PMID: 16008504
- PMCID: PMC1175822
- DOI: 10.1371/journal.pbio.0030273
New insights into metabolic properties of marine bacteria encoding proteorhodopsins
Abstract
Proteorhodopsin phototrophy was recently discovered in oceanic surface waters. In an effort to characterize uncultured proteorhodopsin-exploiting bacteria, large-insert bacterial artificial chromosome (BAC) libraries from the Mediterranean Sea and Red Sea were analyzed. Fifty-five BACs carried diverse proteorhodopsin genes, and we confirmed the function of five. We calculate that proteorhodopsin-exploiting bacteria account for 13% of microorganisms in the photic zone. We further show that some proteorhodopsin-containing bacteria possess a retinal biosynthetic pathway and a reverse sulfite reductase operon, employed by prokaryotes oxidizing sulfur compounds. Thus, these novel phototrophs are an unexpectedly large and metabolically diverse component of the marine microbial surface water.
Figures
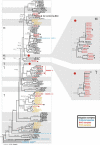
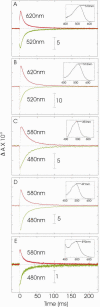
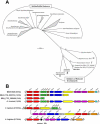

References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

