Shutoff of RNA polymerase II transcription by poliovirus involves 3C protease-mediated cleavage of the TATA-binding protein at an alternative site: incomplete shutoff of transcription interferes with efficient viral replication
- PMID: 16014932
- PMCID: PMC1181600
- DOI: 10.1128/JVI.79.15.9702-9713.2005
Shutoff of RNA polymerase II transcription by poliovirus involves 3C protease-mediated cleavage of the TATA-binding protein at an alternative site: incomplete shutoff of transcription interferes with efficient viral replication
Abstract
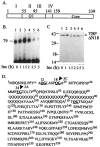
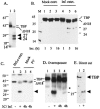
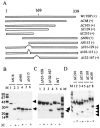
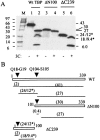
The TATA-binding protein (TBP) plays a crucial role in cellular transcription catalyzed by all three DNA-dependent RNA polymerases. Previous studies have shown that TBP is targeted by the poliovirus (PV)-encoded protease 3C(pro) to bring about shutoff of cellular RNA polymerase II-mediated transcription in PV-infected cells. The processing of the majority of viral precursor proteins by 3C(pro) involves cleavages at glutamine-glycine (Q-G) sites. We present evidence that suggests that the transcriptional inactivation of TBP by 3C(pro) involves cleavage at the glutamine 104-serine 105 (Q104-S105) site of TBP and not at the Q18-G19 site as previously thought. The TBP Q104-S105 cleavage by 3C(pro) is greatly influenced by the presence of an aliphatic amino acid at the P4 position, a hallmark of 3C(pro)-mediated proteolysis. To examine the importance of host cell transcription shutoff in the PV life cycle, stable HeLa cell lines were created that express recombinant TBP resistant to cleavage by the viral proteases, called GG rTBP. Transcription shutoff was significantly impaired and delayed in GG rTBP cells upon infection with poliovirus compared with the cells that express wild-type recombinant TBP (wt rTBP). Infection of GG rTBP cells with poliovirus resulted in small plaques, significantly reduced viral RNA synthesis, and lower viral yields compared to the wt rTBP cell line. These results suggest that a defect in transcription shutoff can lead to inefficient replication of poliovirus in cultured cells.
Figures

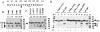
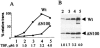

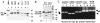

References
-
- Baltimore, D., R. M. Franklin, and J. Callender. 1963. Mengovirus-induced inhibition of host ribonucleic acid and protein. Biochim. Biophys. Acta 76:425-430. - PubMed
-
- Baltimore, D., and R. M. Franklin. 1963. A new ribonucleic acid polymerase appearing after mengovirus infection of L cells. J. Biol. Chem. 238:3395-3400. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

