Bacillus subtilis RecU Holliday-junction resolvase modulates RecA activities
- PMID: 16024744
- PMCID: PMC1176016
- DOI: 10.1093/nar/gki713
Bacillus subtilis RecU Holliday-junction resolvase modulates RecA activities
Abstract
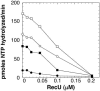
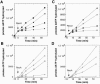
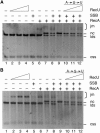
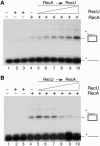
The Bacillus subtilis RecU protein is able to catalyze in vitro DNA strand annealing and Holliday-junction resolution. The interaction between the RecA and RecU proteins, in the presence or absence of a single-stranded binding (SSB) protein, was studied. Substoichiometric amounts of RecU enhanced RecA loading onto single-stranded DNA (ssDNA) and stimulated RecA-catalyzed D-loop formation. However, RecU inhibited the RecA-mediated three-strand exchange reaction and ssDNA-dependent dATP or rATP hydrolysis. The addition of an SSB protein did not reverse the negative effect exerted by RecU on RecA function. Annealing of circular ssDNA and homologous linear 3'-tailed double-stranded DNA by RecU was not affected by the addition of RecA both in the presence and in the absence of SSB. We propose that RecU modulates RecA activities by promoting RecA-catalyzed strand invasion and inhibiting RecA-mediated branch migration, by preventing RecA filament disassembly, and suggest a potential mechanism for the control of resolvasome assembly.
Figures

References
-
- Kowalczykowski S.C. Initiation of genetic recombination and recombination-dependent replication. Trends Biochem. Sci. 2000;25:156–165. - PubMed
-
- Chedin F., Kowalczykowski S.C. A novel family of regulated helicases/nucleases from Gram-positive bacteria: insights into the initiation of DNA recombination. Mol. Microbiol. 2002;43:823–834. - PubMed
-
- Morimatsu K., Kowalczykowski S.C. RecFOR proteins load RecA protein onto gapped DNA to accelerate DNA strand exchange: a universal step of recombinational repair. Mol. Cell. 2003;11:1337–1347. - PubMed
-
- Cox M.M., Goodman M.F., Kreuzer K.N., Sherratt D.J., Sandler S.J., Marians K.J. The importance of repairing stalled replication forks. Nature. 2000;404:37–41. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

