A functional survey of the enhancer activity of conserved non-coding sequences from vertebrate Iroquois cluster gene deserts
- PMID: 16024824
- PMCID: PMC1182218
- DOI: 10.1101/gr.4004805
A functional survey of the enhancer activity of conserved non-coding sequences from vertebrate Iroquois cluster gene deserts
Abstract
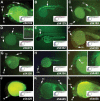
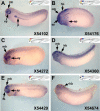
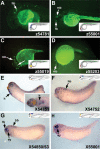
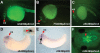
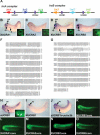
Recent studies of the genome architecture of vertebrates have uncovered two unforeseen aspects of its organization. First, large regions of the genome, called gene deserts, are devoid of protein-coding sequences and have no obvious biological role. Second, comparative genomics has highlighted the existence of an array of highly conserved non-coding regions (HCNRs) in all vertebrates. Most surprisingly, these structural features are strongly associated with genes that have essential functions during development. Among these, the vertebrate Iroquois (Irx) genes stand out on both fronts. Mammalian Irx genes are organized in two clusters (IrxA and IrxB) that span >1 Mb each with no other genes interspersed. Additionally, a large number of HCNRs exist within Irx clusters. We have systematically examined the enhancer activity of HCNRs from the IrxB cluster using transgenic Xenopus and zebrafish embryos. Most of these HCNRs are active in subdomains of endogenous Irx expression, and some are candidates to contain shared enhancers of neighboring genes, which could explain the evolutionary conservation of Irx clusters. Furthermore, HCNRs present in tetrapod IrxB but not in fish may be responsible for novel Irx expression domains that appeared after their divergence. Finally, we have performed a more detailed analysis on two IrxB ultraconserved non-coding regions (UCRs) duplicated in IrxA clusters in similar relative positions. These four regions share a core region highly conserved among all of them and drive expression in similar domains. However, inter-species conserved sequences surrounding the core, specific for each of these UCRs, are able to modulate their expression.
Figures

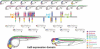
Similar articles
-
Cracking the genome's second code: enhancer detection by combined phylogenetic footprinting and transgenic fish and frog embryos.Methods. 2006 Jul;39(3):212-9. doi: 10.1016/j.ymeth.2005.12.005. Methods. 2006. PMID: 16806968
-
Three enhancer regions regulate gbx2 gene expression in the isthmic region during zebrafish development.Mech Dev. 2006 Dec;123(12):907-24. doi: 10.1016/j.mod.2006.08.007. Epub 2006 Aug 30. Mech Dev. 2006. PMID: 17067785
-
The Xenopus Irx genes are essential for neural patterning and define the border between prethalamus and thalamus through mutual antagonism with the anterior repressors Fezf and Arx.Dev Biol. 2009 May 15;329(2):258-68. doi: 10.1016/j.ydbio.2009.02.028. Epub 2009 Mar 4. Dev Biol. 2009. PMID: 19268445
-
Conservation and diversity in the cis-regulatory networks that integrate information controlling expression of Hoxa2 in hindbrain and cranial neural crest cells in vertebrates.Dev Biol. 2002 Jun 1;246(1):45-56. doi: 10.1006/dbio.2002.0665. Dev Biol. 2002. PMID: 12027433 Review.
-
Distal-less-related homeobox genes of vertebrates: evolution, function, and regulation.Biochem Cell Biol. 2000;78(5):593-601. Biochem Cell Biol. 2000. PMID: 11103950 Review.
Cited by
-
Ancient Pbx-Hox signatures define hundreds of vertebrate developmental enhancers.BMC Genomics. 2011 Dec 30;12:637. doi: 10.1186/1471-2164-12-637. BMC Genomics. 2011. PMID: 22208168 Free PMC article.
-
Ultraconserved Sequences Associated with HoxD Cluster Have Strong Repression Activity.Genome Biol Evol. 2017 Aug 1;9(8):2049-2054. doi: 10.1093/gbe/evx148. Genome Biol Evol. 2017. PMID: 28859354 Free PMC article.
-
Ancient duplicated conserved noncoding elements in vertebrates: a genomic and functional analysis.Genome Res. 2006 Apr;16(4):451-65. doi: 10.1101/gr.4143406. Epub 2006 Mar 13. Genome Res. 2006. PMID: 16533910 Free PMC article.
-
Topologically associating domains are ancient features that coincide with Metazoan clusters of extreme noncoding conservation.Nat Commun. 2017 Sep 5;8(1):441. doi: 10.1038/s41467-017-00524-5. Nat Commun. 2017. PMID: 28874668 Free PMC article.
-
TFCONES: a database of vertebrate transcription factor-encoding genes and their associated conserved noncoding elements.BMC Genomics. 2007 Nov 29;8:441. doi: 10.1186/1471-2164-8-441. BMC Genomics. 2007. PMID: 18045502 Free PMC article.
References
-
- Aparicio, S., Chapman, J., Stupka, E., Putnam, N., Chia, J.M., Dehal, P., Christoffels, A., Rash, S., Hoon, S., Smit, A., et al. 2002. Whole-genome shotgun assembly and analysis of the genome of Fugu rubripes. Science 297: 1301–1310. - PubMed
-
- Bagheri-Fam, S., Ferraz, C., Demaille, J., Scherer, G., and Pfeifer, D. 2001. Comparative genomics of the SOX9 region in human and Fugu rubripes: Conservation of short regulatory sequence elements within large intergenic regions. Genomics 78: 73–82. - PubMed
-
- Bejerano, G., Pheasant, M., Makunin, I., Stephen, S., Kent, W.J., Mattick, J.S., and Haussler, D. 2004. Ultraconserved elements in the human genome. Science 304: 1321–1325. - PubMed
-
- Brenner, S., Elgar, G., Sandford, R., Macrae, A., Venkatesh, B., and Aparicio, S. 1993. Characterization of the pufferfish (Fugu) genome as a compact model vertebrate genome. Nature 366: 265–268. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
