Mutation of the priA gene of Neisseria gonorrhoeae affects DNA transformation and DNA repair
- PMID: 16030229
- PMCID: PMC1196015
- DOI: 10.1128/JB.187.15.5347-5355.2005
Mutation of the priA gene of Neisseria gonorrhoeae affects DNA transformation and DNA repair
Abstract
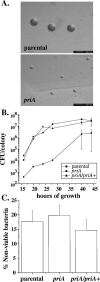
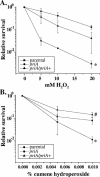
In Escherichia coli, PriA is central to the restart of chromosomal replication when replication fork progression is disrupted and is also involved in homologous recombination and DNA repair. To investigate the role of PriA in recombination and repair in Neisseria gonorrhoeae, we identified, cloned, and insertionally inactivated the gonococcal priA homologue. The priA mutant showed a growth deficiency and decreased DNA repair capability and was completely for deficient in DNA transformation compared to the isogenic parental strain. The priA mutant was also more sensitive to the oxidative damaging agents H2O2 and cumene hydroperoxide compared to the parental strain. These phenotypes were complemented by supplying a functional copy of priA elsewhere in the chromosome. The N. gonorrhoeae priA mutant showed no alteration in the frequency of pilin antigenic variation. We conclude that PriA participates in DNA repair and DNA transformation processes but not in pilin antigenic variation.
Figures



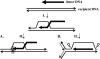
Similar articles
-
Differential roles of homologous recombination pathways in Neisseria gonorrhoeae pilin antigenic variation, DNA transformation and DNA repair.Mol Microbiol. 1998 Nov;30(4):697-710. doi: 10.1046/j.1365-2958.1998.01089.x. Mol Microbiol. 1998. PMID: 10094619
-
Genetic characterization of the nucleotide excision repair system of Neisseria gonorrhoeae.J Bacteriol. 2010 Feb;192(3):665-73. doi: 10.1128/JB.01018-09. Epub 2009 Nov 20. J Bacteriol. 2010. PMID: 19933360 Free PMC article.
-
Characterization of the recD gene of Neisseria gonorrhoeae MS11 and the effect of recD inactivation on pilin variation and DNA transformation.Microbiology (Reading). 1999 Feb;145 ( Pt 2):389-400. doi: 10.1099/13500872-145-2-389. Microbiology (Reading). 1999. PMID: 10075421
-
Dissecting the functional role of PriA protein-catalysed primosome assembly in Escherichia coli DNA replication.Mol Microbiol. 1991 Dec;5(12):2869-73. doi: 10.1111/j.1365-2958.1991.tb01846.x. Mol Microbiol. 1991. PMID: 1667219 Review.
-
Natural transformation of Neisseria gonorrhoeae: from DNA donation to homologous recombination.Mol Microbiol. 2006 Jan;59(2):376-85. doi: 10.1111/j.1365-2958.2005.04964.x. Mol Microbiol. 2006. PMID: 16390436 Review.
Cited by
-
Neisseria gonorrhoeae PBP3 and PBP4 Facilitate NOD1 Agonist Peptidoglycan Fragment Release and Survival in Stationary Phase.Infect Immun. 2019 Jan 24;87(2):e00833-18. doi: 10.1128/IAI.00833-18. Print 2019 Feb. Infect Immun. 2019. PMID: 30510100 Free PMC article.
-
The obligate human pathogen, Neisseria gonorrhoeae, is polyploid.PLoS Biol. 2006 Jun;4(6):e185. doi: 10.1371/journal.pbio.0040185. PLoS Biol. 2006. PMID: 16719561 Free PMC article.
-
Investigating the Role of Helicobacter pylori PriA Protein.Helicobacter. 2016 Aug;21(4):295-304. doi: 10.1111/hel.12283. Epub 2016 Jan 28. Helicobacter. 2016. PMID: 26817518 Free PMC article.
-
ComM is a hexameric helicase that promotes branch migration during natural transformation in diverse Gram-negative species.Nucleic Acids Res. 2018 Jul 6;46(12):6099-6111. doi: 10.1093/nar/gky343. Nucleic Acids Res. 2018. PMID: 29722872 Free PMC article.
-
Replication Restart in Bacteria.J Bacteriol. 2017 Jun 13;199(13):e00102-17. doi: 10.1128/JB.00102-17. Print 2017 Jul 1. J Bacteriol. 2017. PMID: 28320884 Free PMC article. Review.
References
-
- Black, C. G., J. A. M. Fyfe, and J. K. Davies. 1998. Absence of an SOS-like system in Neisseria gonorrhoeae. Gene 208:61-66. - PubMed
-
- Blattner, F. R., G. Plunkett III, C. A. Bloch, N. T. Perna, V. Burland, M. Riley, J. Collado-Vides, J. D. Glasner, C. K. Rode, G. F. Mayhew, J. Gregor, N. W. Davis, H. A. Kirkpatrick, M. A. Goeden, D. J. Rose, B. Mau, and Y. Shao. 1997. The complete genome sequence of Escherichia coli K-12. Science 277:1453-1474. - PubMed
-
- Chen, I., and D. Dubnau. 2003. DNA transport during transformation. Front. Biosci. 8:S544-S556. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

