Arabidopsis SENESCENCE-ASSOCIATED GENE101 stabilizes and signals within an ENHANCED DISEASE SUSCEPTIBILITY1 complex in plant innate immunity
- PMID: 16040633
- PMCID: PMC1197438
- DOI: 10.1105/tpc.105.033910
Arabidopsis SENESCENCE-ASSOCIATED GENE101 stabilizes and signals within an ENHANCED DISEASE SUSCEPTIBILITY1 complex in plant innate immunity
Abstract
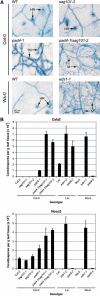
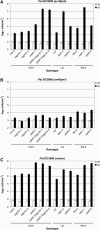
Plant innate immunity against invasive biotrophic pathogens depends on the intracellular defense regulator ENHANCED DISEASE SUSCEPTIBILITY1 (EDS1). We show here that Arabidopsis thaliana EDS1 interacts in vivo with another protein, SENESCENCE-ASSOCIATED GENE101 (SAG101), discovered through a proteomic approach to identify new EDS1 pathway components. Together with PHYTOALEXIN-DEFICIENT4 (PAD4), a known EDS1 interactor, SAG101 contributes intrinsic and indispensable signaling activity to EDS1-dependent resistance. The combined activities of SAG101 and PAD4 are necessary for programmed cell death triggered by the Toll-Interleukin-1 Receptor type of nucleotide binding/leucine-rich repeat immune receptor in response to avirulent pathogen isolates and in restricting the growth of normally virulent pathogens. We further demonstrate by a combination of cell fractionation, coimmunoprecipitation, and fluorescence resonance energy transfer experiments the existence of an EDS1-SAG101 complex inside the nucleus that is molecularly and spatially distinct from EDS1-PAD4 associations in the nucleus and cytoplasm. By contrast, EDS1 homomeric interactions were detected in the cytoplasm but not inside the nucleus. These data, combined with evidence for coregulation between individual EDS1 complexes, suggest that dynamic interactions of EDS1 and its signaling partners in multiple cell compartments are important for plant defense signal relay.
Figures






References
-
- Bhat, R.A., Borst, J.W., Riehl, M., and Thompson, R.D. (2004). Interaction of maize Opaque-2 and the transcriptional co-activators GCN5 and ADA2, in the modulation of transcriptional activity. Plant Mol. Biol. 55, 239–252. - PubMed
-
- Brodersen, P., Petersen, M., Pike, H.M., Olszak, B., Skov, S., Odum, N., Jorgensen, L.B., Brown, R.E., and Mundy, J. (2002). Knockout of Arabidopsis ACCELERATED-CELL-DEATH11 encoding a sphingosine transfer protein causes activation of programmed cell death and defense. Genes Dev. 16, 490–502. - PMC - PubMed
-
- Chandra-Shekara, A.C., Navarre, D., Kachroo, A., Kang, H.G., Klessig, D., and Kachroo, P. (2004). Signaling requirements and role of salicylic acid in HRT- and rrt-mediated resistance to turnip crinkle virus in Arabidopsis. Plant J. 40, 647–659. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

