Involvement of ethylene in stress-induced expression of the TLC1.1 retrotransposon from Lycopersicon chilense Dun
- PMID: 16040666
- PMCID: PMC1183396
- DOI: 10.1104/pp.105.059766
Involvement of ethylene in stress-induced expression of the TLC1.1 retrotransposon from Lycopersicon chilense Dun
Abstract
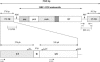
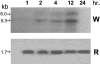
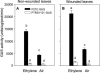
The TLC1 family is one of the four families of long terminal repeat (LTR) retrotransposons identified in the genome of Lycopersicon chilense. Here, we show that this family of retroelements is transcriptionally active and its expression is induced in response to diverse stress conditions such as wounding, protoplast preparation, and high salt concentrations. Several stress-associated signaling molecules, including ethylene, methyl jasmonate, salicylic acid, and 2,4-dichlorophenoxyacetic acid, are capable of inducing TLC1 family expression in vivo. A representative of this family, named TLC1.1, was isolated from a genomic library from L. chilense. Transient expression assays in leaf protoplasts and stably transformed tobacco (Nicotiana tabacum) plants demonstrate that the U3 domain of the 5'-LTR region of this element can drive stress-induced transcriptional activation of the beta-glucuronidase reporter gene. Two 57-bp tandem repeated sequences are found in this region, including an 8-bp motif, ATTTCAAA, previously identified as an ethylene-responsive element box in the promoter region of ethylene-induced genes. Expression analysis of wild-type LTR and single and double ethylene-responsive element box mutants fused to the beta-glucuronidase gene shows that these elements are required for ethylene-responsive gene expression in protoplasts and transgenic plants. We suggest that ethylene-dependent signaling is the main signaling pathway involved in the regulation of the expression of the TLC1.1 element from L. chilense.
Figures






Similar articles
-
The promoter of the TLC1.1 retrotransposon from Solanum chilense is activated by multiple stress-related signaling molecules.Plant Cell Rep. 2007 Oct;26(10):1861-8. doi: 10.1007/s00299-007-0375-y. Epub 2007 Jun 22. Plant Cell Rep. 2007. PMID: 17583815
-
Isolation and molecular characterization of GmERF7, a soybean ethylene-response factor that increases salt stress tolerance in tobacco.Gene. 2013 Jan 15;513(1):174-83. doi: 10.1016/j.gene.2012.10.018. Epub 2012 Oct 27. Gene. 2013. PMID: 23111158
-
Expression of the tobacco Tnt1 retrotransposon promoter in heterologous species.Plant Mol Biol. 1994 Oct;26(1):393-402. doi: 10.1007/BF00039548. Plant Mol Biol. 1994. PMID: 7948885
-
Stress activation and genomic impact of Tnt1 retrotransposons in Solanaceae.Cytogenet Genome Res. 2005;110(1-4):229-41. doi: 10.1159/000084957. Cytogenet Genome Res. 2005. PMID: 16093677 Review.
-
Functional analysis of the tobacco Tnt1 retrotransposon.Genetica. 1994;93(1-3):181-9. doi: 10.1007/BF01435250. Genetica. 1994. PMID: 7813914 Review.
Cited by
-
Transposable elements: multifunctional players in the plant genome.Front Plant Sci. 2024 Jan 4;14:1330127. doi: 10.3389/fpls.2023.1330127. eCollection 2023. Front Plant Sci. 2024. PMID: 38239225 Free PMC article. Review.
-
Ethylene is involved in root phosphorus remobilization in rice (Oryza sativa) by regulating cell-wall pectin and enhancing phosphate translocation to shoots.Ann Bot. 2016 Oct 1;118(4):645-653. doi: 10.1093/aob/mcw044. Ann Bot. 2016. PMID: 27192711 Free PMC article.
-
Corky, a gypsy-like retrotransposon is differentially transcribed in Quercus suber tissues.BMC Res Notes. 2012 Aug 13;5:432. doi: 10.1186/1756-0500-5-432. BMC Res Notes. 2012. PMID: 22888907 Free PMC article.
-
Moving through the Stressed Genome: Emerging Regulatory Roles for Transposons in Plant Stress Response.Front Plant Sci. 2016 Oct 10;7:1448. doi: 10.3389/fpls.2016.01448. eCollection 2016. Front Plant Sci. 2016. PMID: 27777577 Free PMC article. Review.
-
Retrotransposons in Plant Genomes: Structure, Identification, and Classification through Bioinformatics and Machine Learning.Int J Mol Sci. 2019 Aug 6;20(15):3837. doi: 10.3390/ijms20153837. Int J Mol Sci. 2019. PMID: 31390781 Free PMC article. Review.
References
-
- Araujo PJ, Casacuberta JM, Costa AP, Hashimoto RY, Grandbastien MA, Van Sluys MA (2001) Retrolyc1 subfamilies by different U3 LTR regulatory regions in the Lycopersicon genus. Mol Gen Genet 266: 35–41 - PubMed
-
- Bleecker AB, Kende H (2000) Ethylene: a gaseous signal molecule in plant. Annu Rev Cell Dev Biol 16: 1–18 - PubMed
-
- Boeke JD, Corces VG (1989) Transcription and reverse transcription of retrotransposon. Annu Rev Microbiol 43: 403–434 - PubMed
-
- Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72: 248–254 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

