The multifunctional Staphylococcus aureus autolysin aaa mediates adherence to immobilized fibrinogen and fibronectin
- PMID: 16040992
- PMCID: PMC1201280
- DOI: 10.1128/IAI.73.8.4793-4802.2005
The multifunctional Staphylococcus aureus autolysin aaa mediates adherence to immobilized fibrinogen and fibronectin
Abstract
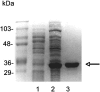
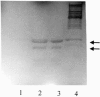
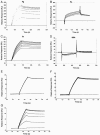
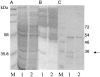
Staphylococci can cause a wide spectrum of infections, including endocarditis, osteomyelitis, and sepsis, which is reflected by the numerous virulence factors they produce, among them a recently identified new class of adhesins, namely, the multifunctional autolysins/adhesins. Here we report the identification and molecular characterization of Aaa, a novel autolysin/adhesin from Staphylococcus aureus. The gene encoding Aaa was cloned from the clinical isolate Staphylococcus aureus 4074. DNA sequence analysis revealed that aaa encodes a deduced protein of 334 amino acids with a predicted molecular mass of 35.8 kDa. Aaa contains three N-terminal repetitive sequences that comprise features of a peptidoglycan-binding domain, the LysM domain. The expression of aaa by Escherichia coli and its subsequent characterization revealed that Aaa possesses bacteriolytic activity as well as adhesive properties, such as binding to extracellular matrix proteins. Real-time biomolecular interaction analysis demonstrated that the interaction of Aaa with fibrinogen, fibronectin, and vitronectin is dose dependent and saturable and occurs with a high affinity. Furthermore, we demonstrate that Aaa binds to the Aalpha and Bbeta chains of fragment D of fibrinogen. Immunofluorescence microscopy revealed that Aaa is located at the cell surface. Finally, an aaa knockout mutant showed reduced adherence to surface-adsorbed fibrinogen and fibronectin, strongly suggesting a role for Aaa in the colonization of host factor-coated polymer surfaces and/or host tissue.
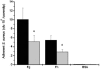
Figures



Similar articles
-
Characterization of the modular design of the autolysin/adhesin Aaa from Staphylococcus aureus.PLoS One. 2012;7(6):e40353. doi: 10.1371/journal.pone.0040353. Epub 2012 Jun 29. PLoS One. 2012. PMID: 22768285 Free PMC article.
-
Identification and characterization of a novel autolysin (Aae) with adhesive properties from Staphylococcus epidermidis.Microbiology (Reading). 2003 Oct;149(Pt 10):2769-2778. doi: 10.1099/mic.0.26527-0. Microbiology (Reading). 2003. PMID: 14523110
-
Functional analysis of a murine monoclonal antibody against the repetitive region of the fibronectin-binding adhesins fibronectin-binding protein A and fibronectin-binding protein B from Staphylococcus aureus.FEBS J. 2010 Nov;277(21):4490-505. doi: 10.1111/j.1742-4658.2010.07835.x. Epub 2010 Sep 28. FEBS J. 2010. PMID: 20875085
-
Sticky connections: extracellular matrix protein recognition and integrin-mediated cellular invasion by Staphylococcus aureus.Curr Opin Microbiol. 2006 Feb;9(1):5-11. doi: 10.1016/j.mib.2005.12.002. Epub 2006 Jan 6. Curr Opin Microbiol. 2006. PMID: 16406780 Review.
-
Structural and functional role of Staphylococcus aureus surface components recognizing adhesive matrix molecules of the host.Future Microbiol. 2009 Dec;4(10):1337-52. doi: 10.2217/fmb.09.102. Future Microbiol. 2009. PMID: 19995192 Review.
Cited by
-
Enterococcus faecium biofilm formation: identification of major autolysin AtlAEfm, associated Acm surface localization, and AtlAEfm-independent extracellular DNA Release.mBio. 2013 Apr 16;4(2):e00154. doi: 10.1128/mBio.00154-13. mBio. 2013. PMID: 23592262 Free PMC article.
-
Identification of antigenic components of Staphylococcus epidermidis expressed during human infection.Infect Immun. 2006 Aug;74(8):4644-54. doi: 10.1128/IAI.00521-06. Infect Immun. 2006. PMID: 16861652 Free PMC article.
-
Modulating activity of vancomycin and daptomycin on the expression of autolysis cell-wall turnover and membrane charge genes in hVISA and VISA strains.PLoS One. 2012;7(1):e29573. doi: 10.1371/journal.pone.0029573. Epub 2012 Jan 9. PLoS One. 2012. PMID: 22253738 Free PMC article.
-
Moonlighting in Rickettsiales: Expanding Virulence Landscape.Trop Med Infect Dis. 2022 Feb 19;7(2):32. doi: 10.3390/tropicalmed7020032. Trop Med Infect Dis. 2022. PMID: 35202227 Free PMC article. Review.
-
Intracellular proteins moonlighting as bacterial adhesion factors.AIMS Microbiol. 2018 May 31;4(2):362-376. doi: 10.3934/microbiol.2018.2.362. eCollection 2018. AIMS Microbiol. 2018. PMID: 31294221 Free PMC article. Review.
References
-
- Allignet, J., P. England, I. Old, and N. El Solh. 2002. Several regions of the repeat domain of the Staphylococcus caprae autolysin, AtlC, are involved in fibronectin binding. FEMS Microbiol. Lett. 213:193-197. - PubMed
-
- Baba, T., F. Takeuchi, M. Kuroda, H. Yuzawa, K. Aoki, A. Oguchi, Y. Nagai, N. Iwama, K. Asano, T. Naimi, H. Kuroda, L. Cui, K. Yamamoto, and K. Hiramatsu. 2002. Genome and virulence determinants of high virulence community-acquired MRSA. Lancet 359:1819-1827. - PubMed
-
- Bateman, A., and M. Bycroft. 2000. The structure of a LysM domain from E. coli membrane-bound lytic murein transglycosylase D (MltD). J. Mol. Biol. 299:1113-1119. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

