High-betweenness proteins in the yeast protein interaction network
- PMID: 16046814
- PMCID: PMC1184047
- DOI: 10.1155/JBB.2005.96
High-betweenness proteins in the yeast protein interaction network
Abstract
Structural features found in biomolecular networks that are absent in random networks produced by simple algorithms can provide insight into the function and evolution of cell regulatory networks. Here we analyze "betweenness" of network nodes, a graph theoretical centrality measure, in the yeast protein interaction network. Proteins that have high betweenness, but low connectivity (degree), were found to be abundant in the yeast proteome. This finding is not explained by algorithms proposed to explain the scale-free property of protein interaction networks, where low-connectivity proteins also have low betweenness. These data suggest the existence of some modular organization of the network, and that the high-betweenness, low-connectivity proteins may act as important links between these modules. We found that proteins with high betweenness are more likely to be essential and that evolutionary age of proteins is positively correlated with betweenness. By comparing different models of genome evolution that generate scale-free networks, we show that rewiring of interactions via mutation is an important factor in the production of such proteins. The evolutionary and functional significance of these observations are discussed.
Figures
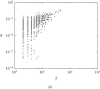

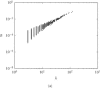



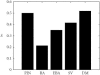
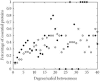
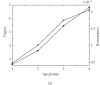

References
-
- Xenarios I, Eisenberg D. Protein interaction databases. Curr Opin Biotechnol. 2001;12(4):334–339. - PubMed
-
- Strogatz S.H. Exploring complex networks. Nature. 2001;410(6825):268–276. - PubMed
-
- Erdos P, Renyi A. On random graphs I. Publicationes Mathematicae. 1959;6:290–297.
-
- Jeong H, Mason S.P, Barabasi A.L, Oltvai Z.N. Lethality and centrality in protein networks. Nature. 2001;411(6833):41–42. - PubMed
-
- Barabasi A.L, Albert R. Emergence of scaling in random networks. Science. 1999;286(5439):509–512. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

