Origin of plasmid-mediated quinolone resistance determinant QnrA
- PMID: 16048974
- PMCID: PMC1196254
- DOI: 10.1128/AAC.49.8.3523-3525.2005
Origin of plasmid-mediated quinolone resistance determinant QnrA
Abstract
Plasmid-mediated resistance to quinolones is increasingly reported in studies of Enterobacteriaceae. Using a PCR-based strategy, a series of gram-negative species were screened for qnrA-like genes. Shewanella algae, an environmental species from marine and fresh water, was identified as its reservoir. This is a one of the very few examples of progenitor identification of an acquired antibiotic resistance gene.
Figures
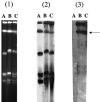
References
-
- Beaber, J. W., B. Hochhut, and M. K. Waldor. 2004. SOS response promotes horizontal dissemination of antibiotic resistance genes. Nature 427:72-74. - PubMed
-
- Brink, A. J., A. Van Straten, and A. J. Rensburg. 1995. Shewanella (Pseudomonas) putrefaciens bacteremia. Clin. Infect. Dis. 20:1327-1332. - PubMed
-
- Fonnesbech-Vogel, B., K. Jorgensen, H. Christensen, J. Elmerdhahl-Olsen, and L. Gram. 1997. Differentiation of Shewanella putrefaciens and Shewanella alga on the basis of whole-cell protein profiles, ribotyping, phenotypic characterization, and 16S rRNA gene sequence analysis. Appl. Environ. Microbiol. 63:2189-2199. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

