Large-scale structural analysis of the core promoter in mammalian and plant genomes
- PMID: 16049029
- PMCID: PMC1181242
- DOI: 10.1093/nar/gki737
Large-scale structural analysis of the core promoter in mammalian and plant genomes
Abstract
DNA encodes at least two independent levels of functional information. The first level is for encoding proteins and sequence targets for DNA-binding factors, while the second one is contained in the physical and structural properties of the DNA molecule itself. Although the physical and structural properties are ultimately determined by the nucleotide sequence itself, the cell exploits these properties in a way in which the sequence itself plays no role other than to support or facilitate certain spatial structures. In this work, we focus on these structural properties, comparing them between different organisms and assessing their ability to describe the core promoter. We prove the existence of distinct types of core promoters, based on a clustering of their structural profiles. These results indicate that the structural profiles are much conserved within plants (Arabidopsis and rice) and animals (human and mouse), but differ considerably between plants and animals. Furthermore, we demonstrate that these structural profiles can be an alternative way of describing the core promoter, in addition to more classical motif or IUPAC-based approaches. Using the structural profiles as discriminatory elements to separate promoter regions from non-promoter regions, reliable models can be built to identify core-promoter regions using a strictly computational approach.
Figures



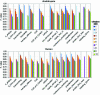

Similar articles
-
Genome-scale computational analysis of DNA curvature and repeats in Arabidopsis and rice uncovers plant-specific genomic properties.BMC Genomics. 2011 May 6;12:214. doi: 10.1186/1471-2164-12-214. BMC Genomics. 2011. PMID: 21548945 Free PMC article.
-
Identification of plant promoter constituents by analysis of local distribution of short sequences.BMC Genomics. 2007 Mar 8;8:67. doi: 10.1186/1471-2164-8-67. BMC Genomics. 2007. PMID: 17346352 Free PMC article.
-
Prediction-based approaches to characterize bidirectional promoters in the mammalian genome.BMC Genomics. 2008;9 Suppl 1(Suppl 1):S2. doi: 10.1186/1471-2164-9-S1-S2. BMC Genomics. 2008. PMID: 18366609 Free PMC article.
-
Computational approaches to identify promoters and cis-regulatory elements in plant genomes.Plant Physiol. 2003 Jul;132(3):1162-76. doi: 10.1104/pp.102.017715. Plant Physiol. 2003. PMID: 12857799 Free PMC article. Review.
-
How can we deliver the large plant genomes? Strategies and perspectives.Curr Opin Plant Biol. 2002 Apr;5(2):173-7. doi: 10.1016/s1369-5266(02)00235-2. Curr Opin Plant Biol. 2002. PMID: 11856615 Review.
Cited by
-
iNuc-PhysChem: a sequence-based predictor for identifying nucleosomes via physicochemical properties.PLoS One. 2012;7(10):e47843. doi: 10.1371/journal.pone.0047843. Epub 2012 Oct 29. PLoS One. 2012. PMID: 23144709 Free PMC article.
-
Unveiling DNA structural features of promoters associated with various types of TSSs in prokaryotic transcriptomes and their role in gene expression.DNA Res. 2017 Feb 1;24(1):25-35. doi: 10.1093/dnares/dsw045. DNA Res. 2017. PMID: 27803028 Free PMC article.
-
Rule-based knowledge acquisition method for promoter prediction in human and Drosophila species.ScientificWorldJournal. 2014;2014:327306. doi: 10.1155/2014/327306. Epub 2014 Jan 29. ScientificWorldJournal. 2014. PMID: 24955394 Free PMC article.
-
DNA structural features of eukaryotic TATA-containing and TATA-less promoters.FEBS Open Bio. 2017 Feb 16;7(3):324-334. doi: 10.1002/2211-5463.12166. eCollection 2017 Mar. FEBS Open Bio. 2017. PMID: 28286728 Free PMC article.
-
Identification of core promoter modules in Drosophila and their application in accurate transcription start site prediction.Nucleic Acids Res. 2006;34(20):5943-50. doi: 10.1093/nar/gkl608. Epub 2006 Oct 26. Nucleic Acids Res. 2006. PMID: 17068082 Free PMC article.
References
-
- Sinden R.R. DNA: Structure and Function. San Diego, CA: Academic press; 1994.
-
- Pedersen A.G., Baldi P., Chauvin Y., Brunak S. DNA structure in human RNA polymerase II promoters. J. Mol. Biol. 1998;281:663–673. - PubMed
-
- Perez-Martin J., de Lorenzo V. Clues and consequences of DNA bending in transcription. Annu. Rev. Microbiol. 1997;51:593–628. - PubMed
-
- Lu X.J., Shakked Z., Olson W.K. A-form conformational motifs in ligand-bound DNA structures. J. Mol. Biol. 2000;300:819–840. - PubMed

