Biological validation of differentially expressed genes in chronic lymphocytic leukemia identified by applying multiple statistical methods to oligonucleotide microarrays
- PMID: 16049305
- PMCID: PMC1867538
- DOI: 10.1016/s1525-1578(10)60562-4
Biological validation of differentially expressed genes in chronic lymphocytic leukemia identified by applying multiple statistical methods to oligonucleotide microarrays
Abstract
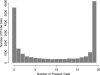
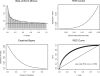
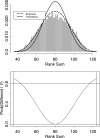
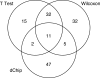
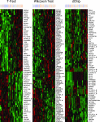
Oligonucleotide microarrays are a powerful tool for profiling the expression levels of thousands of genes. Different statistical methods for identifying differentially expressed genes can yield different results. To our knowledge, no experimental test has been performed to decide which method best identifies genes that are truly differentially expressed. We applied three statistical methods (dChip, t-test on log-transformed data, and Wilcoxon test) to identify differentially expressed genes in previously untreated patients with chronic lymphocytic leukemia (CLL). We used a training set of Affymetrix Hu133A microarray data from 11 patients with unmutated immunoglobulin (Ig) heavy chain variable region (VH) genes and 8 patients with mutated Ig VH genes. Differential expression was validated using semiquantitative real-time polymerase chain reaction assays and by validating models to predict the somatic mutation status of an independent test set of nine CLL samples. The methods identified 144 genes that were differentially expressed between cases of CLL with unmutated compared with mutated Ig VH genes. Eighty genes were identified by Wilcoxon test, 60 by t-test, and 65 by dChip, but only 11 were identified by all three methods. Greater agreement was found between the t-test and the Wilcoxon test. Differential expression was validated by semiquantitative real-time polymerase chain reaction assays for 83% of individual genes, regardless of the statistical method. However, the Wilcoxon test gave the most accurate predictions on new samples, and dChip, the least accurate. We found that all three methods were equally good for finding differentially expressed genes, but they found different genes. The genes selected by the nonparametric Wilcoxon test are the most robust for predicting the status of new cases. A comprehensive list of all differentially expressed genes can only be obtained by combining the results of multiple statistical tests.
Figures
Similar articles
-
Different gene expression in immunoglobulin-mutated and immunoglobulin-unmutated forms of chronic lymphocytic leukemia.Cancer Genet Cytogenet. 2004 Aug;153(1):69-72. doi: 10.1016/j.cancergencyto.2003.12.016. Cancer Genet Cytogenet. 2004. PMID: 15325098
-
ZAP-70 expression identifies a chronic lymphocytic leukemia subtype with unmutated immunoglobulin genes, inferior clinical outcome, and distinct gene expression profile.Blood. 2003 Jun 15;101(12):4944-51. doi: 10.1182/blood-2002-10-3306. Epub 2003 Feb 20. Blood. 2003. PMID: 12595313
-
TWIST2 demonstrates differential methylation in immunoglobulin variable heavy chain mutated and unmutated chronic lymphocytic leukemia.J Clin Oncol. 2005 Jun 10;23(17):3877-85. doi: 10.1200/JCO.2005.02.196. Epub 2005 Apr 4. J Clin Oncol. 2005. PMID: 15809452
-
What do somatic hypermutation and class switch recombination teach us about chronic lymphocytic leukaemia pathogenesis?Curr Top Microbiol Immunol. 2005;294:71-89. doi: 10.1007/3-540-29933-5_5. Curr Top Microbiol Immunol. 2005. PMID: 16323428 Review.
-
The immunoglobulin genes and chronic lymphocytic leukemia (CLL).Ups J Med Sci. 2005;110(2):97-113. doi: 10.3109/2000-1967-075. Ups J Med Sci. 2005. PMID: 16075892 Review.
Cited by
-
Experimental validation of methods for differential gene expression analysis and sample pooling in RNA-seq.BMC Genomics. 2015 Jul 25;16(1):548. doi: 10.1186/s12864-015-1767-y. BMC Genomics. 2015. PMID: 26208977 Free PMC article.
-
A two-gene signature, SKI and SLAMF1, predicts time-to-treatment in previously untreated patients with chronic lymphocytic leukemia.PLoS One. 2011;6(12):e28277. doi: 10.1371/journal.pone.0028277. Epub 2011 Dec 14. PLoS One. 2011. PMID: 22194822 Free PMC article.
-
Identification and validation of biomarkers of IgV(H) mutation status in chronic lymphocytic leukemia using microfluidics quantitative real-time polymerase chain reaction technology.J Mol Diagn. 2007 Sep;9(4):546-55. doi: 10.2353/jmoldx.2007.070001. Epub 2007 Aug 9. J Mol Diagn. 2007. PMID: 17690214 Free PMC article.
-
Optimizing high dimensional gene expression studies for immune response following smallpox vaccination using Taqman® low density immune arrays.J Immunol Methods. 2011 Mar 7;366(1-2):69-78. doi: 10.1016/j.jim.2011.01.011. Epub 2011 Jan 28. J Immunol Methods. 2011. PMID: 21277306 Free PMC article.
References
-
- Damle RN, Wasil T, Fais F, Ghiotto F, Valetto A, Allen SL, Buchbinder A, Budman D, Dittmar K, Kolitz J, Lichtman SM, Schulman P, Vinciguerra VP, Rai KR, Ferrarini M, Chiorazzi N. Ig V gene mutation status and CD38 expression as novel prognostic indicators in chronic lymphocytic leukemia. Blood. 1999;94:1840–1847. - PubMed
-
- Hamblin TJ, Davis Z, Gardiner A, Oscier DG, Stevenson FK. Unmutated Ig V(H) genes are associated with a more aggressive form of chronic lymphocytic leukemia. Blood. 1999;15:1848–1854. - PubMed
-
- McCarthy H, Wierda WG, Barron LL, Cromwell CC, Wang J, Coombes KR, Rangel R, Elenitoba-Johnson KSJ, Keating MJ, Abruzzo LV. High expression of activation-induced cytidine deaminase (AID) and splice variants is a distinctive feature of poor prognosis chronic lymphocytic leukemia. Blood. 2003;101:4903–4908. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

