A family of mammalian E3 ubiquitin ligases that contain the UBR box motif and recognize N-degrons
- PMID: 16055722
- PMCID: PMC1190250
- DOI: 10.1128/MCB.25.16.7120-7136.2005
A family of mammalian E3 ubiquitin ligases that contain the UBR box motif and recognize N-degrons
Abstract
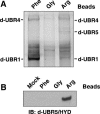
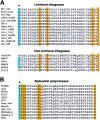
A subset of proteins targeted by the N-end rule pathway bear degradation signals called N-degrons, whose determinants include destabilizing N-terminal residues. Our previous work identified mouse UBR1 and UBR2 as E3 ubiquitin ligases that recognize N-degrons. Such E3s are called N-recognins. We report here that while double-mutant UBR1(-/-) UBR2(-/-) mice die as early embryos, the rescued UBR1(-/-) UBR2(-/-) fibroblasts still retain the N-end rule pathway, albeit of lower activity than that of wild-type fibroblasts. An affinity assay for proteins that bind to destabilizing N-terminal residues has identified, in addition to UBR1 and UBR2, a huge (570 kDa) mouse protein, termed UBR4, and also the 300-kDa UBR5, a previously characterized mammalian E3 known as EDD/hHYD. UBR1, UBR2, UBR4, and UBR5 shared a approximately 70-amino-acid zinc finger-like domain termed the UBR box. The mammalian genome encodes at least seven UBR box-containing proteins, which we propose to call UBR1 to UBR7. UBR1(-/-) UBR2(-/-) fibroblasts that have been made deficient in UBR4 as well (through RNA interference) were significantly impaired in the degradation of N-end rule substrates such as the Sindbis virus RNA polymerase nsP4 (bearing N-terminal Tyr) and the human immunodeficiency virus type 1 integrase (bearing N-terminal Phe). Our results establish the UBR box family as a unique class of E3 proteins that recognize N-degrons or structurally related determinants for ubiquitin-dependent proteolysis and perhaps other processes as well.
Figures






References
-
- Aravind, L., L. M. Iyer, and E. V. Koonin. 2003. Scores of RINGS but no PHDs in ubiquitin signaling. Cell Cycle 2:123-126. - PubMed
-
- Bachmair, A., D. Finley, and A. Varshavsky. 1986. In vivo half-life of a protein is a function of its amino-terminal residue. Science 234:179-186. - PubMed
-
- Bachmair, A., and A. Varshavsky. 1989. The degradation signal in a short-lived protein. Cell 56:1019-1032. - PubMed
-
- Bai, C., P. Sen, K. Hofmann, L. Ma, M. Goebl, J. W. Harper, and S. J. Elledge. 1996. SKP1 connects cell cycle regulators to the ubiquitin proteolysis machinery through a novel motif, the F-box. Cell 86:263-274. - PubMed
-
- Baker, R. T., and A. Varshavsky. 1995. Yeast N-terminal amidase. A new enzyme and component of the N-end rule pathway. J. Biol. Chem. 270:12065-12074. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
