Expressed sequence tags from the oomycete fish pathogen Saprolegnia parasitica reveal putative virulence factors
- PMID: 16076392
- PMCID: PMC1192801
- DOI: 10.1186/1471-2180-5-46
Expressed sequence tags from the oomycete fish pathogen Saprolegnia parasitica reveal putative virulence factors
Abstract
Background: The oomycete Saprolegnia parasitica is one of the most economically important fish pathogens. There is a dramatic recrudescence of Saprolegnia infections in aquaculture since the use of the toxic organic dye malachite green was banned in 2002. Little is known about the molecular mechanisms underlying pathogenicity in S. parasitica and other animal pathogenic oomycetes. In this study we used a genomics approach to gain a first insight into the transcriptome of S. parasitica.
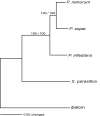
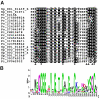
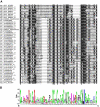
Results: We generated 1510 expressed sequence tags (ESTs) from a mycelial cDNA library of S. parasitica. A total of 1279 consensus sequences corresponding to 525944 base pairs were assembled. About half of the unigenes showed similarities to known protein sequences or motifs. The S. parasitica sequences tended to be relatively divergent from Phytophthora sequences. Based on the sequence alignments of 18 conserved proteins, the average amino acid identity between S. parasitica and three Phytophthora species was 77% compared to 93% within Phytophthora. Several S. parasitica cDNAs, such as those with similarity to fungal type I cellulose binding domain proteins, PAN/Apple module proteins, glycosyl hydrolases, proteases, as well as serine and cysteine protease inhibitors, were predicted to encode secreted proteins that could function in virulence. Some of these cDNAs were more similar to fungal proteins than to other eukaryotic proteins confirming that oomycetes and fungi share some virulence components despite their evolutionary distance
Conclusion: We provide a first glimpse into the gene content of S. parasitica, a reemerging oomycete fish pathogen. These resources will greatly accelerate research on this important pathogen. The data is available online through the Oomycete Genomics Database.
Figures

References
-
- Oomycete Genomics Database (OGD) http://www.oomycete.org
-
- Neish GA, Hughes GC. Fungal diseases of fishes, Book 6. Neptune, New Jersey, USA, T.W.F. Publications; 1980.
-
- Willoughby LG, Pickering AD. Viable Saprolegniaceae spores on the epidermis of the salmonid fish Salmo trutta and Salmo alpinus. Trans Br Mycol Soc. 1977;68:91–95.
-
- Bly JE, Lawson LA, Abdel-Aziz ES, Clem LW. Channel catfish, Ictalurus punctatus, immunity to Saprolegnia sp. J Appl Aquacult. 1994;3:35–50.
-
- Hatai K, Hoshiai GI. Pathogenicity of Saprolegnia parasitica Coker. In: Mueller GJ, editor. Salmon Saprolegniasis. Portland, Oregon, U.S. Department of Energy, Bonneville Power Administration; 1994. pp. 87–98.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

