The pnhA gene of Pasteurella multocida encodes a dinucleoside oligophosphate pyrophosphatase member of the Nudix hydrolase superfamily
- PMID: 16077129
- PMCID: PMC1196092
- DOI: 10.1128/JB.187.16.5809-5817.2005
The pnhA gene of Pasteurella multocida encodes a dinucleoside oligophosphate pyrophosphatase member of the Nudix hydrolase superfamily
Abstract
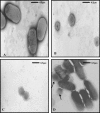
The pnhA gene of Pasteurella multocida encodes PnhA, which is a member of the Nudix hydrolase subfamily of dinucleoside oligophosphate pyrophosphatases. PnhA hydrolyzes diadenosine tetra-, penta-, and hexaphosphates with a preference for diadenosine pentaphosphate, from which it forms ATP and ADP. PnhA requires a divalent metal cation, Mg(2+) or Mn(2+), and prefers an alkaline pH of 8 for optimal activity. A P. multocida strain that lacked a functional pnhA gene, ACP13, was constructed to further characterize the function of PnhA. The cellular size of ACP13 was found to be 60% less than that of wild-type P. multocida, but the growth rate of ACP13 and its sensitivity to heat shock conditions were similar to those of the wild type, and the wild-type cell size was restored in the presence of a functional pnhA gene. Wild-type and ACP13 strains were tested for virulence by using the chicken embryo lethality model, and ACP13 was found to be up to 1,000-fold less virulent than the wild-type strain. This is the first study to use an animal model in assessing the virulence of a bacterial strain that lacked a dinucleoside oligophosphate pyrophosphatase and suggests that the pyrophosphatase PnhA, catalyzing the hydrolysis of diadenosine pentaphosphates, may also play a role in facilitating P. multocida pathogenicity in the host.
Figures


Similar articles
-
The gene ygdP, associated with the invasiveness of Escherichia coli K1, designates a Nudix hydrolase, Orf176, active on adenosine (5')-pentaphospho-(5')-adenosine (Ap5A).J Biol Chem. 2001 Oct 12;276(41):37834-8. doi: 10.1074/jbc.M107032200. Epub 2001 Jul 30. J Biol Chem. 2001. PMID: 11479323
-
Cloning and characterization of AtNUDT13, a novel mitochondrial Arabidopsis thaliana Nudix hydrolase specific for long-chain diadenosine polyphosphates.FEBS J. 2007 Sep;274(18):4877-85. doi: 10.1111/j.1742-4658.2007.06009.x. FEBS J. 2007. PMID: 17824959
-
Characterization of the Mn2+-stimulated (di)adenosine polyphosphate hydrolase encoded by the Deinococcus radiodurans DR2356 nudix gene.Arch Microbiol. 2006 Nov;186(5):415-24. doi: 10.1007/s00203-006-0155-z. Epub 2006 Aug 10. Arch Microbiol. 2006. PMID: 16900379
-
Structures and mechanisms of Nudix hydrolases.Arch Biochem Biophys. 2005 Jan 1;433(1):129-43. doi: 10.1016/j.abb.2004.08.017. Arch Biochem Biophys. 2005. PMID: 15581572 Review.
-
[Pasteurella: insights into the virulence determinants of a heterogenous bacterial type].Berl Munch Tierarztl Wochenschr. 2004 Sep-Oct;117(9-10):367-86. Berl Munch Tierarztl Wochenschr. 2004. PMID: 15495927 Review. German.
Cited by
-
Substrate ambiguity among the nudix hydrolases: biologically significant, evolutionary remnant, or both?Cell Mol Life Sci. 2013 Feb;70(3):373-85. doi: 10.1007/s00018-012-1210-3. Epub 2012 Nov 27. Cell Mol Life Sci. 2013. PMID: 23184251 Free PMC article.
-
The evolution of function within the Nudix homology clan.Proteins. 2017 May;85(5):775-811. doi: 10.1002/prot.25223. Epub 2017 Mar 16. Proteins. 2017. PMID: 27936487 Free PMC article.
-
NrtR Regulates the Type III Secretion System Through cAMP/Vfr Pathway in Pseudomonas aeruginosa.Front Microbiol. 2019 Jan 30;10:85. doi: 10.3389/fmicb.2019.00085. eCollection 2019. Front Microbiol. 2019. PMID: 30761117 Free PMC article.
-
Substrate specificity characterization for eight putative nudix hydrolases. Evaluation of criteria for substrate identification within the Nudix family.Proteins. 2016 Dec;84(12):1810-1822. doi: 10.1002/prot.25163. Epub 2016 Oct 1. Proteins. 2016. PMID: 27618147 Free PMC article.
-
Intruders below the radar: molecular pathogenesis of Bartonella spp.Clin Microbiol Rev. 2012 Jan;25(1):42-78. doi: 10.1128/CMR.05009-11. Clin Microbiol Rev. 2012. PMID: 22232371 Free PMC article. Review.
References
-
- Aguilar, J. S., R. Reyes, A. C. Asensio, S. Oakin, P. Rotllan, and R. Miledi. 2001. Ectoenzymatic breakdown of diadenosine polyphosphates by Xenopus laevis oocytes. Eur. J. Biochem. 268:1289-1297. - PubMed
-
- Ames, B. N., and D. T. Dubin. 1960. The role of polyamines in the neutralization of bacteriophage deoxyribonucleic acid. J. Biol. Chem. 235:769-775. - PubMed
-
- Ausubel, F. M., R. Brent, R. E. Kingston, D. D. Moore, J. G. Seidman, J. A. Smith, and K. Struhl (ed.). 1989. Current protocols in molecular biology. Wiley, New York, N.Y.
-
- Badger, J. L., C. A. Wass, and K. S. Kim. 2000. Identification of Escherichia coli K1 genes contributing to human brain microvascular endothelial cell invasion by differential fluorescence induction. Mol. Microbiol. 36:174-182. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

