Multiple bromodomain genes are involved in restricting the spread of heterochromatic silencing at the Saccharomyces cerevisiae HMR-tRNA boundary
- PMID: 16079223
- PMCID: PMC1456849
- DOI: 10.1534/genetics.105.046938
Multiple bromodomain genes are involved in restricting the spread of heterochromatic silencing at the Saccharomyces cerevisiae HMR-tRNA boundary
Abstract
The transfer RNA gene downstream from the HMR locus in S. cerevisiae functions as part of a boundary (barrier) element that restricts the spread of heterochromatic gene silencing into the downstream region of chromosome III. A genetic screen for identifying additional genes that, when mutated, allow inappropriate spreading of silencing from HMR through the tRNA gene was performed. YTA7, a gene containing bromodomain and ATPase homologies, was identified multiple times. Previously, others had shown that the bromodomain protein Bdf1p functions to restrict silencing at yeast euchromatin-heterochromatin boundaries; therefore we deleted nonessential bromodomain-containing genes to test their effects on heterochromatin spreading. Deletion of RSC2, coding for a component of the RSC chromatin-remodeling complex, resulted in a significant spread of silencing at HMR. Since the bromodomain of YTA7 lacks a key tyrosine residue shown to be important for acetyllysine binding in other bromodomains, we confirmed that a GST-Yta7p bromodomain fusion was capable of binding to histones in vitro. Epistasis analysis suggests that YTA7 and the HMR-tRNA function independently to restrict the spread of silencing, while RSC2 may function through the tRNA element. Our results suggest that multiple bromodomain proteins are involved in restricting the propagation of heterochromatin at HMR.
Figures
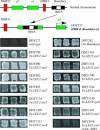



Similar articles
-
A novel role for histone chaperones CAF-1 and Rtt106p in heterochromatin silencing.EMBO J. 2007 May 2;26(9):2274-83. doi: 10.1038/sj.emboj.7601670. Epub 2007 Apr 5. EMBO J. 2007. PMID: 17410207 Free PMC article.
-
The budding yeast protein Chl1p has a role in transcriptional silencing, rDNA recombination, and aging.Biochem Biophys Res Commun. 2005 Nov 11;337(1):167-72. doi: 10.1016/j.bbrc.2005.09.034. Biochem Biophys Res Commun. 2005. PMID: 16182251
-
Cell-cycle control of the establishment of mating-type silencing in S. cerevisiae.Genes Dev. 2002 Nov 15;16(22):2935-45. doi: 10.1101/gad.764102. Genes Dev. 2002. PMID: 12435634 Free PMC article.
-
Genome-wide patterns of histone modifications in yeast.Nat Rev Mol Cell Biol. 2006 Sep;7(9):657-66. doi: 10.1038/nrm1986. Epub 2006 Aug 16. Nat Rev Mol Cell Biol. 2006. PMID: 16912715 Review.
-
Chromatin structure and the regulation of gene expression: the lessons of PEV in Drosophila.Adv Genet. 2008;61:1-43. doi: 10.1016/S0065-2660(07)00001-6. Adv Genet. 2008. PMID: 18282501 Review.
Cited by
-
TFIIIC binding sites function as both heterochromatin barriers and chromatin insulators in Saccharomyces cerevisiae.Eukaryot Cell. 2008 Dec;7(12):2078-86. doi: 10.1128/EC.00128-08. Epub 2008 Oct 10. Eukaryot Cell. 2008. PMID: 18849469 Free PMC article.
-
Dot1 binding induces chromatin rearrangements by histone methylation-dependent and -independent mechanisms.Epigenetics Chromatin. 2011 Feb 3;4(1):2. doi: 10.1186/1756-8935-4-2. Epigenetics Chromatin. 2011. PMID: 21291527 Free PMC article.
-
Direct regulation of nucleosome density by the conserved AAA-ATPase Yta7.Proc Natl Acad Sci U S A. 2011 Dec 6;108(49):E1302-11. doi: 10.1073/pnas.1116819108. Epub 2011 Nov 9. Proc Natl Acad Sci U S A. 2011. PMID: 22074782 Free PMC article.
-
Transcription independent insulation at TFIIIC-dependent insulators.Genetics. 2009 Sep;183(1):131-48. doi: 10.1534/genetics.109.106203. Epub 2009 Jul 13. Genetics. 2009. PMID: 19596900 Free PMC article.
-
Direct interplay among histones, histone chaperones, and a chromatin boundary protein in the control of histone gene expression.Mol Cell Biol. 2012 Nov;32(21):4337-49. doi: 10.1128/MCB.00871-12. Epub 2012 Aug 20. Mol Cell Biol. 2012. PMID: 22907759 Free PMC article.
References
-
- Burns, N., B. Grimwade, P. B. Ross-Macdonald, E. Y. Choi, K. Finberg et al., 1994. Large-scale analysis of gene expression, protein localization, and gene disruption in Saccharomyces cerevisiae. Genes Dev. 8: 1087–1105. - PubMed
-
- Cairns, B. R., Y. Lorch, Y. Li, M. Zhang, L. Lacomis et al., 1996. RSC, an essential, abundant chromatin-remodeling complex. Cell 87: 1249–1260. - PubMed
-
- Carmen, A. A., L. Milne and M. Grunstein, 2002. Acetylation of the yeast histone H4 N terminus regulates its binding to heterochromatin protein SIR3. J. Biol. Chem. 277: 4778–4781. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

