Distribution of cryptosporidium genotypes in storm event water samples from three watersheds in New York
- PMID: 16085835
- PMCID: PMC1183313
- DOI: 10.1128/AEM.71.8.4446-4454.2005
Distribution of cryptosporidium genotypes in storm event water samples from three watersheds in New York
Abstract
To assess the source and public health significance of Cryptosporidium oocyst contamination in storm runoff, a PCR-restriction fragment length polymorphism technique based on the small-subunit rRNA gene was used in the analysis of 94 storm water samples collected from the Malcolm Brook and N5 stream basins in New York over a 3-year period. The distribution of Cryptosporidium in this study was compared with the data obtained from 27 storm water samples from the Ashokan Brook in a previous study. These three watersheds represented different levels of human activity. Among the total of 121 samples analyzed from the three watersheds, 107 were PCR positive, 101 of which (94.4%) were linked to animal sources. In addition, C. hominis (W14) was detected in six samples collected from the Malcolm Brook over a 2-week period. Altogether, 22 Cryptosporidium species or genotypes were found in storm water samples from these three watersheds, only 11 of which could be attributed to known species/groups of animals. Several Cryptosporidium spp. were commonly found in these three watersheds, including the W1 genotype from an unknown animal source, the W4 genotype from deer, and the W7 genotype from muskrats. Some genotypes were found only in a particular watershed. Aliquots of 113 samples were also analyzed by the Environmental Protection Agency (EPA) Method 1623; 63 samples (55.7%) were positive for Cryptosporidium by microscopy, and 39 (78%) of the 50 microscopy-negative samples were positive by PCR. Results of this study demonstrate that molecular techniques can complement traditional detection methods by providing information on the source of contamination and the human-infective potential of Cryptosporidium oocysts found in water.
Figures


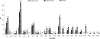


References
-
- Deng, M. Q., and D. O. Cliver. 1999. Improved immunofluorescence assay for detection of Giardia and Cryptosporidium from asymptomatic adult cervine animals. Parasitol. Res. 85:733-736. - PubMed
-
- Faubert, G. M., and Y. Litvinsky. 2000. Natural transmission of Cryptosporidium parvum between dams and calves on a dairy farm. J. Parasitol. 86:495-500. - PubMed
-
- Fayer, R., J. M. Trout, E. J. Lewis, M. Santin, L. Zhou, A. A. Lal, and L. Xiao. 2003. Contamination of Atlantic coast commercial shellfish with Cryptosporidium. Parasitol. Res. 89:141-145. - PubMed
-
- Fox, K. R., and D. A. Lytle. 1996. Milwaukee's crypto outbreak: investigation and recommendations. J. Am. Water Assoc. 88:87-94.
-
- Graczyk, T. K., M. R. Cranfield, and R. Fayer. 1996. Evaluation of commercial enzyme immunoassay (EIA) and immunofluorescent antibody (FA) test kits for detection of Cryptosporidium oocysts of species other than Cryptosporidium parvum. Am. J. Trop. Med. Hyg. 54:274-279. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

