Multiscale modeling of cardiac cellular energetics
- PMID: 16093514
- PMCID: PMC2864600
- DOI: 10.1196/annals.1341.035
Multiscale modeling of cardiac cellular energetics
Abstract
Multiscale modeling is essential to integrating knowledge of human physiology starting from genomics, molecular biology, and the environment through the levels of cells, tissues, and organs all the way to integrated systems behavior. The lowest levels concern biophysical and biochemical events. The higher levels of organization in tissues, organs, and organism are complex, representing the dynamically varying behavior of billions of cells interacting together. Models integrating cellular events into tissue and organ behavior are forced to resort to simplifications to minimize computational complexity, thus reducing the model's ability to respond correctly to dynamic changes in external conditions. Adjustments at protein and gene regulatory levels shortchange the simplified higher-level representations. Our cell primitive is composed of a set of subcellular modules, each defining an intracellular function (action potential, tricarboxylic acid cycle, oxidative phosphorylation, glycolysis, calcium cycling, contraction, etc.), composing what we call the "eternal cell," which assumes that there is neither proteolysis nor protein synthesis. Within the modules are elements describing each particular component (i.e., enzymatic reactions of assorted types, transporters, ionic channels, binding sites, etc.). Cell subregions are stirred tanks, linked by diffusional or transporter-mediated exchange. The modeling uses ordinary differential equations rather than stochastic or partial differential equations. This basic model is regarded as a primitive upon which to build models encompassing gene regulation, signaling, and long-term adaptations in structure and function. During simulation, simpler forms of the model are used, when possible, to reduce computation. However, when this results in error, the more complex and detailed modules and elements need to be employed to improve model realism. The processes of error recognition and of mapping between different levels of model form complexity are challenging but are essential for successful modeling of large-scale systems in reasonable time. Currently there is to this end no established methodology from computational sciences.
Figures

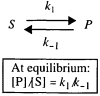
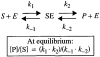
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

