Comparative analysis of expression of histone H2a genes in mouse
- PMID: 16098230
- PMCID: PMC1198228
- DOI: 10.1186/1471-2164-6-108
Comparative analysis of expression of histone H2a genes in mouse
Abstract
Background: At least 18 replication-dependent histone H2a genes are distributed in 3 Hist gene clusters on different chromosomes of the mouse genome. In this analysis we designed specific PCR primers for each histone H2a transcript and studied the expression levels and patterns using quantitative RT-PCR (qRT-PCR). In addition, we compared histone H3 K9 acetylation levels in the promoter regions of H2a genes by ChIP (chromatin immunoprecipitation)--quantitative PCR (qPCR) analysis.
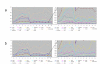
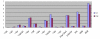
Results: RT-PCR analysis indicated that all 20 histone H2a genes assessed in this study are expressed. The replication-dependent histone H2a genes have different expression levels but similar expression patterns. Among the 20 histone H2a genes, the expression-level of H2afz, a replication-independent gene, was highest, and that of Hist1h2aa, a replication-dependent gene, was lowest. Among 18 replication-dependent H2a genes, the expression level of Hist3h2a was highest. The ChIP-qPCR analysis showed that histone H3 K9 acetylation levels in promoter regions of both H2afz and Hist3h2a are clearly higher than that in the promoter region of Hist1h2aa. The H3 K9 acetylation level in the promoter of Hist1h2aa is similar to that in the gamma-satellite region.
Conclusion: These results strongly suggest that histone H3 K9 acetylation plays a role in the expression of histone genes.
Figures

Similar articles
-
A novel replication-independent histone H2a gene in mouse.BMC Genet. 2005 Feb 19;6:10. doi: 10.1186/1471-2156-6-10. BMC Genet. 2005. PMID: 15720718 Free PMC article.
-
Naturally occurring antisense RNA of histone H2a in mouse cultured cell lines.BMC Genet. 2005 May 14;6:23. doi: 10.1186/1471-2156-6-23. BMC Genet. 2005. PMID: 15892893 Free PMC article.
-
The malaria parasite Plasmodium falciparum histones: organization, expression, and acetylation.Gene. 2006 Mar 15;369:53-65. doi: 10.1016/j.gene.2005.10.022. Epub 2006 Jan 10. Gene. 2006. PMID: 16410041
-
Site-specific analysis of histone methylation and acetylation.Methods Mol Biol. 2004;287:99-120. doi: 10.1385/1-59259-828-5:099. Methods Mol Biol. 2004. PMID: 15273407 Review.
-
Epigenetic control of ovarian function: the emerging role of histone modifications.Mol Cell Endocrinol. 2005 Nov 24;243(1-2):12-8. doi: 10.1016/j.mce.2005.09.005. Epub 2005 Oct 10. Mol Cell Endocrinol. 2005. PMID: 16219412 Review.
Cited by
-
Transcriptional inhibition after irradiation occurs preferentially at highly expressed genes in a manner dependent on cell cycle progression.bioRxiv [Preprint]. 2024 Jun 11:2023.11.20.567799. doi: 10.1101/2023.11.20.567799. bioRxiv. 2024. Update in: Elife. 2024 Oct 11;13:RP94001. doi: 10.7554/eLife.94001. PMID: 38045243 Free PMC article. Updated. Preprint.
-
RUNX1 regulates site specificity of DNA demethylation by recruitment of DNA demethylation machineries in hematopoietic cells.Blood Adv. 2017 Sep 6;1(20):1699-1711. doi: 10.1182/bloodadvances.2017005710. eCollection 2017 Sep 12. Blood Adv. 2017. PMID: 29296817 Free PMC article.
-
Distribution of introns in fungal histone genes.PLoS One. 2011 Jan 27;6(1):e16548. doi: 10.1371/journal.pone.0016548. PLoS One. 2011. PMID: 21304581 Free PMC article.
-
A screening system to identify transcription factors that induce binding site-directed DNA demethylation.Epigenetics Chromatin. 2017 Dec 8;10(1):60. doi: 10.1186/s13072-017-0169-6. Epigenetics Chromatin. 2017. PMID: 29221486 Free PMC article.
-
Genome-wide analysis of abnormal H3K9 acetylation in cloned mice.PLoS One. 2008 Apr 9;3(4):e1905. doi: 10.1371/journal.pone.0001905. PLoS One. 2008. PMID: 18398451 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

