Differential expression of the Arabidopsis cytochrome c genes Cytc-1 and Cytc-2. Evidence for the involvement of TCP-domain protein-binding elements in anther- and meristem-specific expression of the Cytc-1 gene
- PMID: 16113211
- PMCID: PMC1203360
- DOI: 10.1104/pp.105.065920
Differential expression of the Arabidopsis cytochrome c genes Cytc-1 and Cytc-2. Evidence for the involvement of TCP-domain protein-binding elements in anther- and meristem-specific expression of the Cytc-1 gene
Abstract
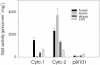
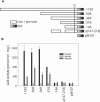
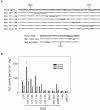
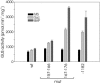
The promoters of the Arabidopsis (Arabidopsis thaliana) cytochrome c genes, Cytc-1 and Cytc-2, were analyzed using plants transformed with fusions to the beta-glucuronidase coding sequence. Histochemical staining of plants indicated that the Cytc-1 promoter directs preferential expression in root and shoot meristems and in anthers. In turn, plants transformed with the Cytc-2 promoter fusions showed preferential expression in vascular tissues of cotyledons, leaves, roots, and hypocotyls, and also in anthers. Quantitative measurements in extracts prepared from different organs suggested that expression of Cytc-1 is higher in flowers, while that of Cytc-2 is higher in leaves. The analysis of a set of deletions and site-directed mutants of the Cytc-1 promoter indicated that a segment located between -147 and -156 from the translation start site is required for expression and that site II elements (TGGGCC/T) located in this region, coupled with a downstream internal telomeric repeat (AAACCCTAA), are responsible for the expression pattern of this gene. Proteins present in cauliflower nuclear extracts, as well as a recombinant protein from the TCP-domain family, were able to specifically bind to the region required for expression. We propose that expression of the Cytc-1 gene is linked to cell proliferation through the elements described above. The fact that closely located site II motifs are present in similar locations in several genes encoding proteins involved in cytochrome c-dependent respiration suggests that these elements may be the target of factors that coordinate the expression of nuclear genes encoding components of this part of the mitochondrial respiratory chain.
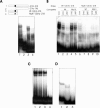
Figures


Similar articles
-
A segment containing a G-box and an ACGT motif confers differential expression characteristics and responses to the Arabidopsis Cytc-2 gene, encoding an isoform of cytochrome c.J Exp Bot. 2009;60(3):829-45. doi: 10.1093/jxb/ern331. Epub 2008 Dec 19. J Exp Bot. 2009. PMID: 19098132
-
The promoter of the Arabidopsis nuclear gene COX5b-1, encoding subunit 5b of the mitochondrial cytochrome c oxidase, directs tissue-specific expression by a combination of positive and negative regulatory elements.J Exp Bot. 2004 Sep;55(405):1997-2004. doi: 10.1093/jxb/erh223. Epub 2004 Jul 30. J Exp Bot. 2004. PMID: 15286148
-
Common sets of promoter elements determine the expression characteristics of three Arabidopsis genes encoding isoforms of mitochondrial cytochrome c oxidase subunit 6b.Plant Cell Physiol. 2009 Jul;50(7):1393-9. doi: 10.1093/pcp/pcp080. Epub 2009 Jun 3. Plant Cell Physiol. 2009. PMID: 19493962
-
The multiple functions of cytochrome c and their regulation in life and death decisions of the mammalian cell: From respiration to apoptosis.Mitochondrion. 2011 May;11(3):369-81. doi: 10.1016/j.mito.2011.01.010. Epub 2011 Feb 4. Mitochondrion. 2011. PMID: 21296189 Free PMC article. Review.
-
Anticancer Nanoparticle Carriers of the Proapoptotic Protein Cytochrome c.Pharmaceutics. 2025 Feb 26;17(3):305. doi: 10.3390/pharmaceutics17030305. Pharmaceutics. 2025. PMID: 40142969 Free PMC article. Review.
Cited by
-
Tissue-specific transcriptomic profiling of Sorghum propinquum using a rice genome array.PLoS One. 2013;8(3):e60202. doi: 10.1371/journal.pone.0060202. Epub 2013 Mar 25. PLoS One. 2013. PMID: 23536906 Free PMC article.
-
Redox modulation of plant developmental regulators from the class I TCP transcription factor family.Plant Physiol. 2013 Jul;162(3):1434-47. doi: 10.1104/pp.113.216416. Epub 2013 May 17. Plant Physiol. 2013. PMID: 23686421 Free PMC article.
-
Characterization of Arabidopsis thaliana genes encoding functional homologues of the yeast metal chaperone Cox19p, involved in cytochrome c oxidase biogenesis.Plant Mol Biol. 2007 Oct;65(3):343-55. doi: 10.1007/s11103-007-9224-1. Epub 2007 Aug 22. Plant Mol Biol. 2007. PMID: 17712601
-
Delta subclass HD-Zip proteins and a B-3 AP2/ERF transcription factor interact with promoter elements required for expression of the Arabidopsis cytochrome c oxidase 5b-1 gene.Plant Mol Biol. 2012 Sep;80(2):157-67. doi: 10.1007/s11103-012-9935-9. Epub 2012 Jun 6. Plant Mol Biol. 2012. PMID: 22669746
-
Characterization of PISTILLATA-like Genes and Their Promoters from the Distyly Fagopyrum esculentum.Plants (Basel). 2022 Apr 12;11(8):1047. doi: 10.3390/plants11081047. Plants (Basel). 2022. PMID: 35448776 Free PMC article.
References
-
- Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16: 735–743 - PubMed
-
- Cubas P, Lauter N, Doebley J, Coen E (1999) The TCP domain: a motif found in proteins regulating plant growth and development. Plant J 18: 215–222 - PubMed
-
- Curi GC, Welchen E, Chan RL, Gonzalez DH (2003) Nuclear and mitochondrial genes encoding cytochrome c oxidase subunits respond differently to the same metabolic factors. Plant Physiol Biochem 41: 689–693
-
- Elorza A, León G, Gómez I, Mouras A, Holuigue L, Araya A, Jordana X (2004) Nuclear SDH2-1 and SDH2-2 genes, encoding the iron-sulfur subunit of mitochondrial complex II in Arabidopsis, have distinct cell-specific expression patterns and promoter activities. Plant Physiol 136: 4072–4087 - PMC - PubMed
-
- Felitti SA, Chan RL, Gago G, Valle EM, Gonzalez DH (1997) Expression of sunflower cytochrome c mRNA is tissue-specific and controlled by nitrate and light. Physiol Plant 99: 342–347
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

