The most infectious prion protein particles
- PMID: 16148934
- PMCID: PMC1513539
- DOI: 10.1038/nature03989
The most infectious prion protein particles
Abstract
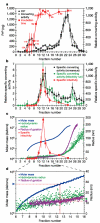
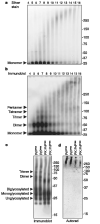
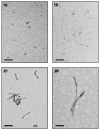
Neurodegenerative diseases such as Alzheimer's, Parkinson's and the transmissible spongiform encephalopathies (TSEs) are characterized by abnormal protein deposits, often with large amyloid fibrils. However, questions have arisen as to whether such fibrils or smaller subfibrillar oligomers are the prime causes of disease. Abnormal deposits in TSEs are rich in PrP(res), a protease-resistant form of the PrP protein with the ability to convert the normal, protease-sensitive form of the protein (PrP(sen)) into PrP(res) (ref. 3). TSEs can be transmitted between organisms by an enigmatic agent (prion) that contains PrP(res) (refs 4 and 5). To evaluate systematically the relationship between infectivity, converting activity and the size of various PrP(res)-containing aggregates, PrP(res) was partially disaggregated, fractionated by size and analysed by light scattering and non-denaturing gel electrophoresis. Our analyses revealed that with respect to PrP content, infectivity and converting activity peaked markedly in 17-27-nm (300-600 kDa) particles, whereas these activities were substantially lower in large fibrils and virtually absent in oligomers of < or =5 PrP molecules. These results suggest that non-fibrillar particles, with masses equivalent to 14-28 PrP molecules, are the most efficient initiators of TSE disease.
Figures
References
-
- Caughey B, Lansbury PT. Protofibrils, pores, fibrils, and neurodegeneration: separating the responsible protein aggregates from the innocent bystanders. Annu. Rev. Neurosci. 2003;26:267–298. - PubMed
-
- Kayed R, et al. Permeabilization of lipid bilayers is a common conformation-dependent activity of soluble amyloid oligomers in protein misfolding diseases. J. Biol. Chem. 2004;279:46363–46366. - PubMed
-
- Kocisko DA, et al. Cell-free formation of protease-resistant prion protein. Nature. 1994;370:471–474. - PubMed
-
- Silveira JR, Caughey B, Baron GS. Prion protein and the molecular features of transmissible spongiform encephalopathy agents. Curr. Top. Microbiol. Immunol. 2004;284:1–50. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

