FOXO1a acts as a selective tumor suppressor in alveolar rhabdomyosarcoma
- PMID: 16157701
- PMCID: PMC2171446
- DOI: 10.1083/jcb.200501040
FOXO1a acts as a selective tumor suppressor in alveolar rhabdomyosarcoma
Retraction in
-
FOXO1a acts as a selective tumor suppressor in alveolar rhabdomyosarcoma.J Cell Biol. 2007 May 7;177(3):563. doi: 10.1083/jcb.20050104020070420r. J Cell Biol. 2007. PMID: 17485494 Free PMC article. No abstract available.
Abstract
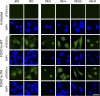
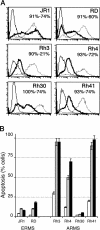
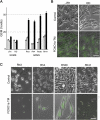
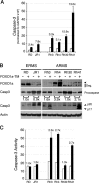
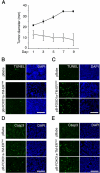
Rhabdomyosarcoma (RMS), the most common pediatric soft-tissue sarcoma, has two major histological subtypes: embryonal RMS (ERMS), which has a favorable prognosis, and alveolar RMS (ARMS), which has a poor outcome. Although both forms of RMS express muscle cell-specific markers, only ARMS cells express PAX3-FOXO1a or PAX7-FOXO1a chimeric proteins. In mice, Pax3 and Pax7 play key roles in muscle cell development and differentiation, and FoxO1a regulates myoblast differentiation and fusion; thus, the aberrant regulation of these proteins may contribute to the development of ARMS. In this paper, we report that FOXO1a is not expressed in primary ARMS tumors or ARMS-derived tumor cell lines and that restoration of FOXO1a expression in ARMS cells is sufficient to induce cell cycle arrest and apoptosis. Strikingly, the effects of FOXO1a are selective, as enforced expression of FOXO1a in ERMS-derived tumor cell lines had no effect. Furthermore, FOXO1a induced apoptosis in ARMS by directly activating the transcription of caspase-3. We conclude that FOXO1a is a potent and specific tumor suppressor in ARMS, suggesting that agents that restore or augment FOXO1a activity may be effective as ARMS therapeutics.
Figures

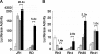



Comment in
-
Findings of misconduct in science/research misconduct.NIH Guide Grants Contracts (Bethesda). 2011 Jun 17:NOT-OD-11-084. NIH Guide Grants Contracts (Bethesda). 2011. PMID: 21714172 Free PMC article. No abstract available.
-
Findings of Misconduct in Science/Research Misconduct.Fed Regist. 2011 Jun 9;76(111):33763-33764. Fed Regist. 2011. PMID: 27737203 Free PMC article. No abstract available.
References
-
- Accili, D., and K.C. Arden. 2004. FoxOs at the crossroads of cellular metabolism, differentiation, and transformation. Cell. 117:421–426. - PubMed
-
- Ausubel, F.M., R. Brent, R.E. Kingston, D.D. Moore. J.G. Seidman, J.A. Smith, and K. Struhl. 2001. Current Protocols in Molecular Biology. John Wiley and Sons, Inc., Somerset, NJ. 1512 pp.
-
- Barr, F.G. 2001. Gene fusions involving PAX and FOX family members in alveolar rhabdomyosarcoma. Oncogene. 20:5736–5746. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

